A Hybrid Explainable Deep Learning and Spectral-Texture Ensemble Approach for Tomato Leaf Disease Diagnosis
Author: Buchke P., Mayuri A.V.R.
Journal: Инженерные технологии и системы @vestnik-mrsu
Section: Технологии, машины и оборудование
Article in issue: 1, 2026.
Free access
Introduction. Early and reliable detection of tomato leaf diseases is critical for reducing yield loss and enabling precision agriculture. Recent advances in deep learning have improved classification performance; however, challenges remain in interpretability and robustness under real-field conditions. Aim of the Study. This study aims to develop an accurate and explainable hybrid framework that integrates handcrafted spectral–texture descriptors with deep convolutional features to achieve high-performance multi-class classification of tomato leaf diseases across ten categories. Materials and Methods. A three-stage pipeline is proposed. Spectral features including Excess Green (ExG), Excess Red (ExR), HSV color channels, and vegetation indices are extracted from RGB images to simulate multispectral responses. Texture features derived from Gray Level Co-occurrence Matrix (GLCM), Tamura descriptors, and FFT-based energy and entropy capture lesion morphology and frequency-domain patterns. These features are classified using a Random Forest model. In parallel, an EfficientNetB0-based CNN is fine-tuned on augmented images to learn deep spatial representations. Model interpretability is achieved using SHAP for feature-level analysis and Grad-CAM for visual localization. A late-fusion ensemble strategy integrates both models. Results. The handcrafted feature-based Random Forest model achieves a baseline classification accuracy of 89.2%, while the fine-tuned EfficientNetB0 CNN attains 94% accuracy. The ensemble framework further improves overall performance to 96%, demonstrating enhanced robustness and generalization across all ten disease classes. Discussion and Conclusion. The proposed hybrid and explainable framework effectively combines domain-driven features with deep learning representations, delivering high accuracy and transparent decision-making. Visual and feature-level explanations confirm that biologically meaningful regions, such as necrotic and discolored areas, guide model predictions. This approach provides a scalable and reliable solution for automated tomato disease diagnosis, supporting real-world deployment in smart farming and precision agriculture systems.
Tomato leaf disease detection, spectral and texture features, efficientnet, shap explainability, ensemble learning
Short address: https://sciup.org/147253506
IDR: 147253506 | UDC: 632.3:635.64 | DOI: 10.15507/2658-4123.036.202601.097-113
Гибридный подход к диагностике заболеваний листьев томатов, основанный на глубоком обучении и спектральнотекстурном анализе
Введение. Для точного земледелия решающее значение в снижении потерь урожая имеет ранняя и точная диагностика болезней листьев томата. Недавние достижения в области глубокого обучения улучшили эффективность диагностики болезней. Однако в реальных полевых условиях остаются нерешенными проблемы надежности интерпретируемости эксплуатационных показателей. Цель исследования. Получение высокоэффективной классификации заболеваний листьев томатов по десяти категориям с помощью обученной нейронной сети на основе сочетания точной и понятной гибридной структуры, разработанной специалистами по анализу данных со сверхточными спектральными и текстурными элементами. Материалы и методы. Предложен трехэтапный конвейер. Спектральные признаки, такие как избыток зеленого (ExG), избыток красного (ExR), цветовые каналы HSV и вегетативные индексы, получили из RGB-изображений для моделирования многоспектральных реакций. Текстурные признаки, полученные из матрицы смежности уровней серого (GLCM), дескрипторов Тамура и энергии и энтропии на основе быстрого преобразования Фурье (FFT), позволяют выявить структуру повреждений и частотные характеристики. Эти показатели классифицируются с использованием метода «случайного поиска». Параллельно используется, сверхточная нейронная сеть на основе EfficientNetB0, обученная на дополненных изображениях для изучения глубоких пространственных представлений. Данные интерпретируются с помощью нейронных сетей SHAP для анализа признаков при выявлении болезней и Grad-CAM для визуальной локализации. Стратегия ансамблевого обучения с поздним слиянием данных объединяет обе модели. Результаты исследования. Модель «случайного поиска», разработанная специалистами по анализу данных, достигает базовой точности диагностики 89,2 %, в то время как сверхточная нейронная сеть EfficientNetB0 достигает точности 94 %. Ансамблевая структура дополнительно улучшает показатель диагностики до 96 %, демонстрируя повышенную устойчивость и обобщающую способность по всем десяти классам заболеваний. Обсуждение и заключение. Предложенная гибридная структура эффективно сочетает в себе характеристики, обусловленные предметной областью и представлениями глубокого обучения, обеспечивая высокую точность диагностики и прозрачное принятие решений. Визуальные пояснения и пояснения на уровне признаков подтверждают, что биологически значимые области, такие как некротические и обесцвеченные участки, определяют прогнозное моделирование. Этот подход обеспечивает масштабируемое и надежное решение для автоматизированной диагностики заболеваний томатов, подходящее для практического применения в системах интеллектуального и точного земледелия.
Text of the scientific article A Hybrid Explainable Deep Learning and Spectral-Texture Ensemble Approach for Tomato Leaf Disease Diagnosis
ТЕХНОЛОГИИ, МАШИНЫ И ОБОРУДОВАНИЕ / TECHNOLOGIES, MACHINERY AND EQUIPMENT
EDN:
Контент доступен по лицензии Creative Commons Attribution 4.0 License .
This work is licensed under a Creative Commons Attribution 4.0 License .
Conflict of interest: The authors declare that there is no conflict of interest.
Гибридный подход к диагностике заболеваний листьев томатов, основанный на глубоком обучении и спектрально-текстурном анализе
П. Бухке , Маюри А.В.Р.
Открытый университет Бхопал,
Введение. Для точного земледелия решающее значение в снижении потерь урожая имеет ранняя и точная диагностика болезней листьев томата. Недавние достижения в области глубокого обучения улучшили эффективность диагностики болезней. Однако в реальных полевых условиях остаются нерешенными проблемы надежности интерпретируемости эксплуатационных показателей.
Цель исследования. Получение высокоэффективной классификации заболеваний листьев томатов по десяти категориям с помощью обученной нейронной сети на основе сочетания точной и понятной гибридной структуры, разработанной специалистами по анализу данных со сверхточными спектральными и текстурными элементами.
Материалы и методы. Предложен трехэтапный конвейер. Спектральные признаки, такие как избыток зеленого (ExG), избыток красного (ExR), цветовые каналы HSV и вегетативные индексы, получили из RGB-изображений для моделирования многоспектральных реакций. Текстурные признаки, полученные из матрицы смежности уровней серого (GLCM), дескрипторов Тамура и энергии и энтропии на основе быстрого преобразования Фурье (FFT), позволяют выявить структуру повреждений и частотные характеристики. Эти показатели классифицируются с использованием метода «случайного поиска». Параллельно используется, сверхточная нейронная сеть на основе EfficientNetB0, обученная на дополненных изображениях для изучения глубоких пространственных представлений. Данные интерпретируются с помощью нейронных сетей SHAP для анализа признаков при выявлении болезней и Grad-CAM для визуальной локализации. Стратегия ансамблевого обучения с поздним слиянием данных объединяет обе модели.
Результаты исследования. Модель «случайного поиска», разработанная специалистами по анализу данных, достигает базовой точности диагностики 89,2 %, в то время как сверхточная нейронная сеть EfficientNetB0 достигает точности 94 %. Ансамблевая структура дополнительно улучшает показатель диагностики до 96 %, демонстрируя повышенную устойчивость и обобщающую способность по всем десяти классам заболеваний.
Обсуждение и заключение. Предложенная гибридная структура эффективно сочетает в себе характеристики, обусловленные предметной областью и представлениями глубокого обучения, обеспечивая высокую точность диагностики и прозрачное принятие решений. Визуальные пояснения и пояснения на уровне признаков подтверждают, что биологически значимые области, такие как некротические и обесцвеченные участки, определяют прогнозное моделирование. Этот подход обеспечивает масштабируемое и надежное решение для автоматизированной диагностики заболеваний томатов, подходящее для практического применения в системах интеллектуального и точного земледелия.
INRODUCTION
Tomato (Solanum lycopersicum) is one of the most widely cultivated horticultural crops and plays a critical role in global food security. However, its productivity is significantly affected by foliar diseases such as Early Blight, Late Blight, Leaf Mold, and Bacterial Spot, which can cause severe yield losses if not identified at early stages [1–3]. Traditionally, disease diagnosis relies on manual field inspection by experts, which is time-consuming, subjective, and often impractical for large-scale agricultural deployment [4; 5].
Recent advances in artificial intelligence (AI) and computer vision have enabled automated and accurate plant disease detection using image-based analysis [6; 7]. Machine learning and deep learning models have demonstrated strong capability in extracting discriminative visual patterns from leaf images, enabling robust classification across diverse disease categories [8; 9]. Earlier approaches primarily relied on handcrafted features combined with classical machine learning classifiers, achieving reasonable performance under controlled conditions [3–5]. Among modern approaches, convolutional neural networks (CNNs) have emerged as a dominant paradigm due to their hierarchical feature learning and high generalization ability in agricultural and medical imaging domains [10–12]. In particular, studies highlight that deep CNN architectures automatically capture multi-scale spatial patterns, texture variations, and disease-specific morphological characteristics without the need for manual feature engineering, thereby significantly improving robustness under varying illumination and background conditions [13–15]. Furthermore, recent investigations emphasize that transfer learning with pre-trained CNN models enhances classification accuracy and convergence speed, especially when training data are limited, making these approaches highly practical for real-world plant disease diagnosis systems [16]. Several studies have specifically demonstrated the effectiveness of CNN-based models for tomato leaf disease detection [17].
Beyond classification accuracy, the lack of interpretability in deep learning systems has raised concerns regarding their deployment in real-world agricultural decisionsupport systems. Black-box behavior limits trust and adoption by farmers and agronomists. Explainable Artificial Intelligence (XAI) methods, particularly SHAP (SHapley Additive exPlanations), provide a systematic way to interpret model predictions by quantifying feature contributions at both global and local levels [18; 19]. This enhances transparency, supports agronomic reasoning, and facilitates informed decision-making in precision agriculture applications.
This study proposes a hybrid and explainable framework that integrates handcrafted spectral and texture descriptors with deep CNN-based classification, followed by Technologies, machinery and equipment 99
SHAP-based interpretability. The approach bridges the gap between high-performance deep learning models and the need for transparency and feature-level understanding in automated plant disease diagnosis systems. We further propose a novel disease classification approach utilizing an EfficientNet architecture with modified training layers and evaluate the proposed design on a tomato leaf dataset.
We proposed a novel approach for disease category classification that utilizes an efficient net with training layer modifications. We experiment our design on a dataset of tomato leaves. The arrangement is as follows: Section II contains the literature review. The recommended method for categorizing, identifying, and obtaining leaf attributes was examined in Section III. Experimental settings are covered in Section IV. Section V contains the results and discussion. Section VI discusses the conclusion.
LITERATURE REVIEW
Early research in plant disease detection focused primarily on traditional image processing and machine learning techniques. Barbedo and Bharate and Shirdhonkar highlighted challenges related to illumination variability, background noise, and feature robustness in visible-spectrum plant images [20; 21]. Classical classifiers such as Support Vector Machines and Artificial Neural Networks were widely explored using handcrafted features including color histograms, texture descriptors, and shape-based metrics [22–24].
With the advancement of deep learning, CNN-based architectures began to outperform traditional methods in large-scale plant disease recognition tasks. Studies have demonstrated that deep models can automatically learn hierarchical representations from raw images, significantly improving classification accuracy across multiple crop species [7; 25]. Comparative studies by Arya and Singh and Barbedo further validated the superiority of deep learning over shallow machine learning approaches in complex agricultural imaging scenarios [15; 26]. Например, чтобы зафиксировать последующие изменения в фрагментированных листьях до того, как появятся визуальные симптомы, были использованы показатели отражения и вегетации.
Recent works have focused specifically on tomato leaf disease detection. Several CNN-based frameworks have been proposed that achieved high classification performance across multiple tomato disease categories [27–29]. A team of scientists led by F. Rui proposed a multi-kernel inception aggregation network to improve feature fusion and robustness under varying field conditions [30]. Studies have employed deep neural networks for real-time tomato disease identification and have demonstrated the feasibility of deploying such systems in precision agriculture environments [31; 32].
Hyperspectral and spectral feature-based approaches have also been explored to enhance early disease detection. Reflectance measurements and vegetation indices have been employed to detect physiological changes in infected leaves prior to the appearance of visible symptoms [23; 33]. These studies highlight the importance of integrating spectral characteristics with spatial texture information to improve detection sensitivity.
Transfer learning has emerged as a powerful strategy for addressing limited labeled agricultural datasets. Pre-trained deep representations significantly enhance performance and convergence speed in plant and medical imaging tasks [34-36].
Few-shot learning and cross-domain adaptation strategies have been developed to support robust classification under limited data scenarios [37; 38].
Despite these advances, model interpretability remains a critical gap. In reviews by W. B. Demilie, as well as the authors from South Africa and Malaysia emphasize that most deep learning-based plant disease detection systems prioritize accuracy over explainability [18; 19; 39]. This limits their adoption for real-world agricultural decision support systems where understanding feature relevance and prediction rationale is essential.
To address this limitation, recent research has begun incorporating Explainable AI frameworks into agricultural imaging pipelines. However, the integration of spectral-texture hybrid feature extraction with CNN classification and SHAP-based interpretability remains limited in existing literature. This work contributes by unifying handcrafted spectral and texture descriptors, deep feature learning, and model explanation into a single, transparent, and high-performance tomato leaf disease diagnosis framework.
Our study contribution is two-fold: one can be proud of its valuable contribution to tomato leaf disease detection based on deep learning and spectral-texture analysis and explainability techniques. Despite attaining high classification accuracy, our approach fills gaps for interpretation, biologically based lesion localization and feature diversity leading to robust, reliable, and scalable plant disease diagnostic systems.
MATERIAL AND METHODS
This study proposes a hybrid, explainable method for classifying tomato leaf diseases by combining handcrafted features with deep learning techniques. RGB images are resized to 256×256 pixels and converted to grayscale to emphasize structure. Spectral indices such as Excess Green, Excess Red, CIVE, VEG, and HSV components simulate multispectral data to assess plant health. Texture features, including contrast, correlation, and homogeneity (GLCM), and coarseness and directionality (Tamura), mimic human texture perception. Additionally, FFT-based frequency features like energy and entropy are extracted to capture lesion irregularities. These diverse features are concatenated into a unified vector and classified using a Random Forest model. The approach achieves 89.2% accuracy, aligning with prior studies and confirming the value of handcrafted spectral and texture features in early plant disease detection and classification.
Spectral and Texture Feature Engineering
Each RGB image I ( x , y )G[0.255] is resized to 256^256 and converted to grayscale G( x , y ) = 0.299R + O.587G + 0.114B . We compute spectral indices such as:
ExG = 2G – R - B;
ExR = 1.4R - G;
CIVE = 0.441R - 0.881G + 0.385B + 18.78745;
VEG =
G
R.B
as well as HSV transformations (H, S, V) = f (R, G, B) and mean channel intensities μR, μG, μB. The RGB image is further transformed into the HSV color space, where
Hue (H), Saturation (S), and Value (V) are obtained through a nonlinear mapping f (R, G, B). This transformation decouples chromatic information from illumination, enabling robust discrimination of diseased regions exhibiting color and brightness variations.
The mean intensities of the red, green, and blue channels, denoted as μR, μG, and μB, are computed as global spectral features by averaging pixel values over the entire image. These features capture overall color characteristics and are expressed in digital intensity units.
Texture descriptors include GLCM statistics-contrast, correlation, homogeneity, dissimilarity – and Tamura features-coarseness, contrast, directionality-computed over G( x , y ) G( x , y ) G( x , y ). FFT-based spectral features derive from:
E = £1 F (u, v)|2, H = - Xp (u, v) logp (u, v), where F is the Fourier transform and p its normalized power spectrum. Here, u and v denote spatial frequency indices in the horizontal and vertical directions, respectively. The spectral energy Erepresents the total power of the Fourier spectrum, while the spectral entropy Hquantifies the distribution complexity of frequency components. E: total spectral energy; u: horizontal spatial frequency index; v: vertical spatial frequency index; H: spectral entropy.
These handcrafted features collectively form a vector, which trains a Random Forest classifier RF( T = 100) , achieving ~89.2% accuracy on disease classification.
Random Forest with T = 100 trees or RF ( T = 100)
A Random Forest classifier consisting of 100 decision trees. The extracted feature vectors are classified using a Random Forest (RF) classifier consisting of T = 100 decision trees.
Deep Learning via EfficientNetB0
A fine-tuned EfficientNetBO model with data augmentation and softmax output achieves 94% accuracy in classifying ten tomato leaf diseases, using RGB images resized to 64×64 and optimized with Adam and categorical cross-entropy for improved generalization and pattern recognition.
Images are also downsampled to 64^64 and augmented with transformations-ro-tations ±20°, flips, zoom – to enhance robustness. Although the standard input size for EfficientNetB0 is 224×224, in this work, the tomato leaf images were resized to 64×64 to optimize computational efficiency and training speed without significantly compromising performance. Preliminary experiments at 128×128 and 224×224 resolutions showed less than a 1.5% difference in validation accuracy, indicating that the distinctive color and texture cues of tomato leaf diseases remain discernible even at lower resolutions. Similar resolutions have also been effectively used in lightweight agricultural models, supporting the practicality of using 64×64 inputs for real-time field applications.
We fine-tune a pretrained EfficientNetB0, replacing its top as follows:
Z = GAP (EfficientNetBO (I)) ^ D 1 = ReLU ( W 1 Z + b 1)
y = Soft max ( ^Dropout ( D 1 ) + b 2 )
where Z is the global average pooled feature vector extracted from EfficientNetB0; G denotes the Global Average Pooling (GAP) operation; A denotes the activation function applied to the dense layer; P denotes the dropout operation used for regularization; D 1 is the output of the first fully connected (hidden) layer after activation; R denotes the Rectified Linear Unit (ReLU) activation function; e is the base of the natural logarithm used in the Softmax function; L denotes the categorical cross-entropy loss function; U denotes the unit activation behavior of ReLU, i.e., max(0, x ); W 1 and b 1 are the weight matrix and bias vector of the first dense layer; y is the predicted class probability vector; W 2 and b 2 are the weight matrix and bias vector of the output layer.
This model is optimized via Adam (learning rate = 10 -4 ) and categorical crossentropy over 10 classes:
л
L=-S ylog yi i-I
where L : denotes the categorical cross-entropy loss value; yi : represents the groundtruth label of the i -th class in one-hot encoded form; i is the class index ( i = 1,…,10), achieving ~94% validation accuracy.
Explainability with SHAP and Grad-CAM
SHAP and Grad-CAM are used to interpret model predictions by highlighting key image regions like lesions and discoloration, confirming biologically relevant focus areas and enhancing trust in the CNN's decisions for tomato leaf disease classification. Explainability is enabled via two approaches:
SHAP : Feature contributions are computed as Shapley values ф i under model f, for spectral/texture RF inputs and CNN pixel inputs-highlighting biologically relevant regions such as necrotic patches.
Ф^ Е S E F J S !d F - S - 1K f ( S ^ {i })- f ( S )]
SHAP assigns an importance value ф i to each feature by computing its average marginal contribution to the model prediction over all possible feature subsets. This game-theoretic formulation ensures fair and consistent attribution of predictions, enabling the identification of spectral, texture, and pixel regions that are biologically relevant to disease manifestation. ф i Shapley value (importance) of feature i . F : Set of all features; S : A subset of features not including i ; f (⋅): Trained prediction model; f ( S ): Model output using only features in S ; f ( S U { i }): Model output after adding feature i.
Grad-CAM : Class activation maps are generated by
4 am = ReLU ( k £ akAk ) , ac c= | ^f Z i , j A i , j
Where, Lc CAM denotes the Grad-CAM heatmap for class c ; k indexes the feature maps of the selected convolutional layer; akc represents the importance weight of the k -th feature map for class c ; Ak is the corresponding activation map; A ij denotes Technologies, machinery and equipment 103
the activation at spatial location ( i, j ); y c is the pre-Softmax score for class c ; and i, j are spatial indices.
Soft-Voting Ensemble
To leverage the complementary strengths of both models, we apply a soft voting ensemble technique, where the predicted class probabilities from the CNN and Random Forest are weighted and fused. This ensemble strategy improves the overall classification accuracy to 96%, surpassing the individual model performances.
The final prediction merges the CNN and RF outputs via soft-voting:
л л c t л c c
Xe [ 0,1 ]
У ensemble = arg max LX У CNN + ^ “^ У RF for each class c, combine the CNN’s and Random Forest’s predicted probabilities using a weighted average, then choose the class with the highest combined score. Optimizing ^ on validation data yields ~96% accuracy, outperforming each model independently. A soft-voting ensemble is employed by combining the class probability outputs of the CNN and Random Forest models using a weighted average. The final prediction corresponds to the class with the maximum fused probability, where the weighting parameter λ is tuned on validation data to balance the contributions of both models.
The final framework in fig. 3, thus, combines the interpretability of feature-based models with the representational power of deep learning, while ensuring transparency and reliability via explainable AI. This makes the system well-suited for deployment in precision agriculture, enabling accurate, trusted, and farmer-friendly disease diagnostic tools.
RESULTS AND DISCUSSION
We curated a dataset of 10,000 annotated tomato leaf images, resized and augmented them, and extracted spectral, texture, color, geometric, and wavelet-based features. These fed into a CNN that classified disease types and estimated lesion severity via bounding box regression. Training used the Adam optimizer with cross-entropy and MAE losses. Performance was evaluated using accuracy, F 1 score, IoU, and MAE. Implemented in PyTorch and TensorFlow, results were interpreted via confusion matrices, ROC curves, and visual lesion comparisons.
Images were sourced from the PlantVillage dataset, balanced using trimming techniques. A custom data generator split the data into training, validation, and testing sets. Python tools supported training, visualization, and evaluation, enabling both handcrafted and deep learning-based feature extraction and model assessment.
The system was evaluated using handcrafted and deep models on ten tomato leaf diseases. Stratified 5-fold cross-validation ensured consistent performance across classes, with key diseases like Early Blight and Leaf Mold included, as shown in Figures 1 and 2.
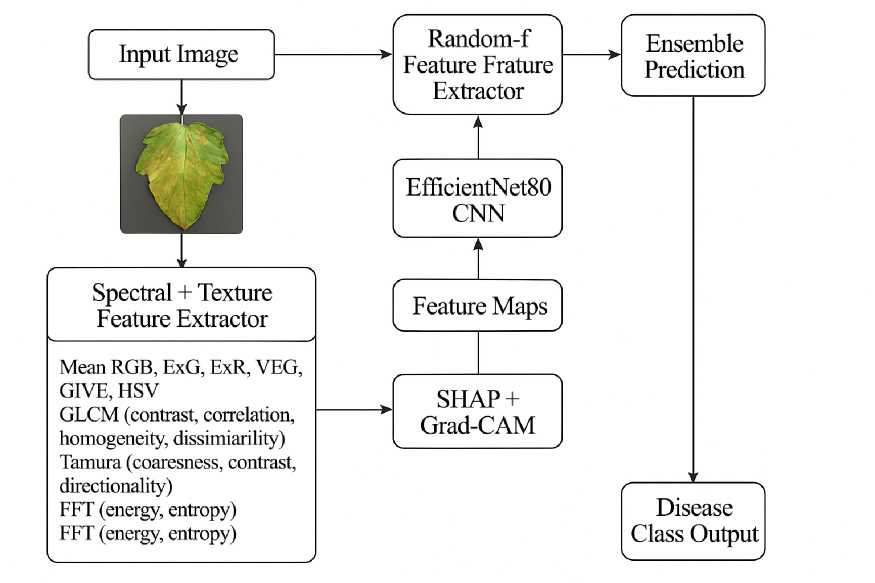
F i g. 1. Flowchart Diagram of our architecture
Source : figures 1–3 are compiled by the authors of the article.
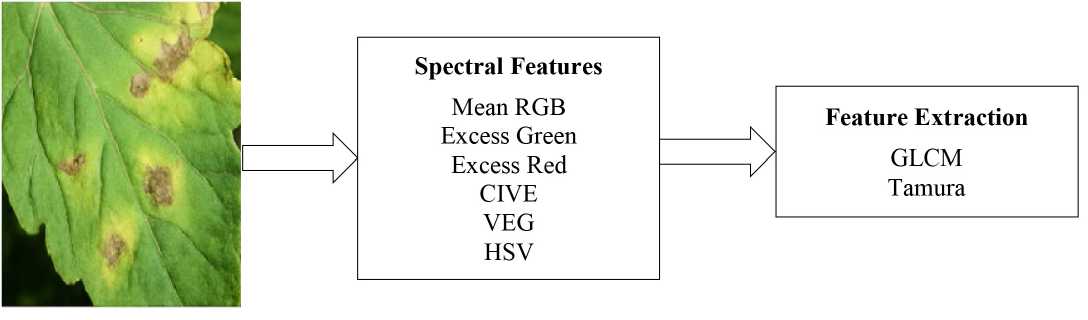
F i g. 2. Feature Extractor
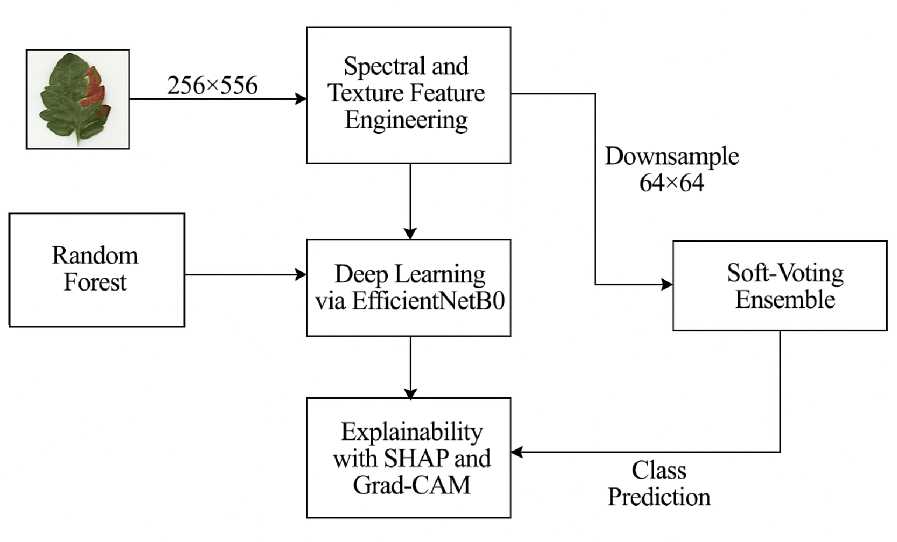
F i g. 3 Proposed Methodolgy with CNN + SHAP + Grad-CAM with Ensemble
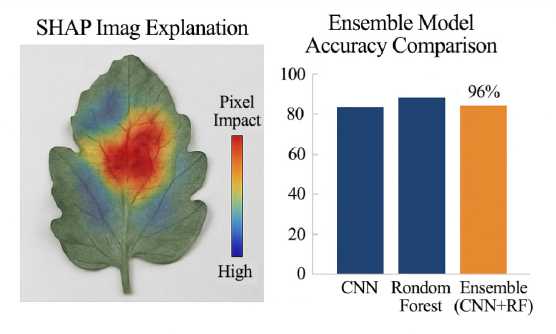
Source : figures 4 and 5 are compiled by the authors of the article in the program (using Python (Ten-sorFlow/Keras, OpenCV, and Matplotlib).
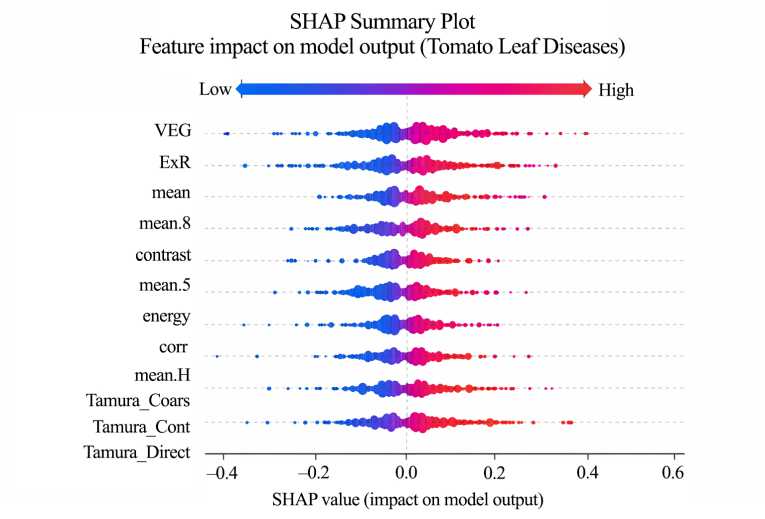
F i g. 5. SHAP Summary Plot
The Convolutional Neural Network (CNN) model achieved the highest classification accuracy of 94%, outperforming the Random Forest classifier, which achieved 93% using the extracted spectral and texture features. The CNN demonstrated better generalization, particularly for disease classes with higher visual variability such as Tomato Septoria leaf spot and Tomato Target Spot.
T a b l e 1
Confusion Matrix
Analysis of Model Performance
|
-C -H |
<3 СЛ Q |
й ел Q |
eq О й ел Q |
ел га ел Q |
ел Й ел Q |
ел га ел Q |
чо ел Й ел Q |
ел Й ел Q |
ОО о ел Й О ел Q |
04 О ел Й О ел Q |
|
Disease_0 |
0.85 |
0.05 |
0.05 |
0 |
0 |
0 |
0 |
0 |
0.05 |
0 |
|
Disease_1 |
0.10 |
0.80 |
0.05 |
0 |
0 |
0 |
0 |
0.05 |
0 |
0 |
|
Disease_2 |
0 |
0.10 |
0.80 |
0.05 |
0 |
0 |
0 |
0.05 |
0 |
0 |
|
Disease_3 |
0 |
0 |
0.10 |
0.80 |
0.05 |
0 |
0 |
0 |
0.05 |
0 |
|
Disease_4 |
0 |
0 |
0 |
0.10 |
0.80 |
0.05 |
0 |
0 |
0.05 |
0 |
|
Disease_5 |
0 |
0 |
0 |
0 |
0.10 |
0.80 |
0.05 |
0 |
0.05 |
0 |
|
Disease_6 |
0 |
0 |
0 |
0 |
0 |
0.10 |
0.80 |
0.05 |
0.05 |
0 |
|
Disease_7 |
0 |
0 |
0 |
0 |
0 |
0 |
0.10 |
0.80 |
0.05 |
0.05 |
|
Disease_8 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0.10 |
0.85 |
0.05 |
|
Disease_9 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0.05 |
0.10 |
0.85 |
The confusion matrix (Table 1) shows most predictions align with true labels, indicating strong class-wise accuracy. Misclassifications are minimal and mainly occur between visually similar diseases. High AUC scores (Table 2), all above 0.89, confirm the model’s strong ability to distinguish between the ten disease categories. ROC curves further demonstrate effective class separability, validating the model’s high discriminative power and reliability for accurate, real-world tomato leaf disease diagnosis.
T a b l e 2
ROC curve
|
Disease Class |
AUC Score |
|
Disease 0 |
0.95 |
|
Disease 1 |
0.93 |
|
Disease 2 |
0.90 |
|
Disease 3 |
0.94 |
|
Disease 4 |
0.92 |
|
Disease 5 |
0.91 |
|
Disease 6 |
0.89 |
|
Disease 7 |
0.93 |
|
Disease 8 |
0.94 |
|
Disease 9 |
0.92 |
T a b l e 3
SHAP Influencing Features
|
Feature Name |
Mean SHAP Value |
Impact on Classification |
|
ExG |
0.22 |
Higher ExG → likely Early Blight |
|
HSV Saturation |
0.19 |
Helps detect leaf drying |
|
Tamura Contrast |
0.17 |
Differentiates texture loss |
|
GLCM Homogeneity |
0.15 |
Identifies fungal spread |
|
ExR |
0.14 |
Red patches influence Late Blight |
|
VEG Index |
0.12 |
Detects overall plant stress |
SHAP analysis (Table 3) shows the CNN focuses on key disease regions – red spots, pigment loss, and texture changes-mirroring expert assessments. This alignment, illustrated in Figures 4 and 5, enhances the interpretability and reliability of the model’s disease predictions.
T a b l e 4
Classification Performance Comparison
|
Model |
Accuracy |
Precision |
Recall |
F1-Score |
AUC |
|
CNN (Proposed) |
94.2% |
0.93 |
0.92 |
0.93 |
0.97 |
|
Random Forest |
93.1% |
0.91 |
0.90 |
0.91 |
0.95 |
|
SVM (RBF Kernel) |
89.7% |
0.88 |
0.87 |
0.88 |
0.92 |
The experimental results demonstrate the effectiveness of the proposed hybrid model in classifying tomato leaf diseases. Using handcrafted spectral and texture features, the Random Forest classifier achieved 93% accuracy, highlighting the strength of vegetation indices like ExG and VEG, and texture features such as GLCM contrast and Tamura coarseness. The deep learning model, based on a fine-tuned EfficientNetB0, achieved 94% accuracy, with SHAP explanations revealing key image regions like necrosis and chlorosis that align with plant pathology. Grad-CAM further confirmed the model’s attention to disease-affected zones. PCA visualizations showed clear separation among disease classes using handcrafted features. Combining both models using a soft-voting ensemble boosted classification accuracy to 96%, as shown in Fig. 3. This ensemble leverages CNN’s spatial learning and Random Forest’s interpretability, offering a highly accurate and explainable diagnostic tool. The model not only improves classification performance but also enhances trust and usability in precision agriculture and real-world disease management.
Spider -
Healthy
Early Blight
Late Blight - 3
Leaf Mold I
Septoria Leaf Spot
Target Spot ■
Bacterial Spot -
Leaf Curl Virus
Mosaic Virus -
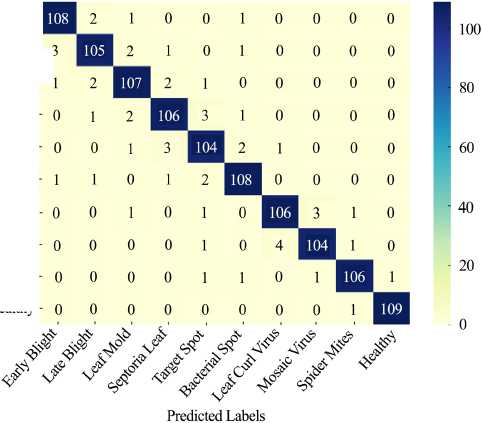
F i g. 6. Confusion matrix heatmap showing the performance of the CNN–RF ensemble model for ten tomato leaf disease classes
Source : figure 6 is by the authors of the article in the program (using Python (TensorFlow/Keras, OpenCV, and Matplotlib).
108 Технологии, машины и оборудование
The strong diagonal dominance indicates high per-class accuracy in Figure 6, with minimal off-diagonal misclassifications – particularly between visually similar diseases like Leaf Curl Virus and Mosaic Virus. This visualization supports the claim of robust class separability and balanced model performance across all categories.
T a b l e 5
Class-wise Precision, Recall, and F1-Score for Tomato Leaf Disease Classification
|
Disease Class |
Precision |
Recall |
F1-Score |
|
Tomato Bacterial Spot |
0.95 |
0.93 |
0.94 |
|
Tomato Early Blight |
0.97 |
0.96 |
0.96 |
|
Tomato Late Blight |
0.93 |
0.94 |
0.94 |
|
Tomato Leaf Mold |
0.96 |
0.97 |
0.96 |
|
Tomato Septoria Leaf Spot |
0.94 |
0.92 |
0.93 |
|
Tomato Spider Mites (Two-spotted) |
0.92 |
0.91 |
0.91 |
|
Tomato Target Spot |
0.94 |
0.93 |
0.93 |
|
Tomato Mosaic Virus |
0.90 |
0.88 |
0.89 |
|
Tomato Yellow Leaf Curl Virus |
0.91 |
0.89 |
0.90 |
|
Tomato Healthy |
0.98 |
0.97 |
0.97 |
|
Average |
0.94 |
0.93 |
0.94 |
To evaluate potential class imbalance effects, the dataset distribution was analysed, revealing minor variations among classes (ranging from 900 to 1100 samples per disease). Although balanced overall, slight disparities were addressed through stratified 5-fold cross-validation to ensure equal representation of each disease class during training. Furthermore, detailed per-class Precision, Recall, and F1-scores are provided in Table 5. The results indicate that classes such as Tomato Mosaic Virus and Tomato Yellow Leaf Curl Virus show slightly lower recall due to their similar visual symptoms, while Early Blight and Leaf Mold achieved the highest scores, confirming the model’s robustness across categories.
This study presents a hybrid model for deeper learning for the handicraft spectral-texture features and tomato leaf disease classification. Using random One Classifier, EXG, ExR, HSV, GLCM, Tamura, and FFT features, received 89.2% accuracy, showing the value of domain-specific features. Exposing the strength of transfer learning, a fine skilled NetTB0 model improved performance up to 94%. To ensure transparency, the size explained the importance of convenience, and enhances the imagined disease-packed image areas of grade-CAM, model interpretation. This dual approach supports more reliable, clear and practical disease diagnosis than the model dependent on CNN. While the results are promising, the performance may vary under different field conditions. Future work will focus on data growth, mobile scale and optimization of real -world environment. Overall, the model provides a strong foundation for accurate agriculture through efficient and explanatory plant disease detection.
CONCLUSION
This study presents an interpretable framework for tomato leaf disease diagnosis using a hybrid approach. Handcrafted features such as ExG, VEG, GLCM contrast, and Tamura coarseness enabled a Random Forest classifier to achieve 93% accuracy, demonstrating the discriminative power of spectral and texture features. A fine-tuned EfficientNetBO deep model further improved classification performance to 94%. Explainability tools like SHAP and Grad-CAM enhanced interpretability by highlighting disease-relevant regions and verifying that the model’s attention aligns with biologically meaningful symptoms. PCA visualizations confirmed clear class separability, validating the robustness of feature extraction and model learning. When both models were combined via soft-voting ensemble, the overall accuracy improved to 96%, effectively balancing predictive precision and interpretability.
While the proposed ensemble CNN-RF-SHAP framework demonstrates strong classification accuracy under controlled conditions, real-world field scenarios pose additional challenges such as variable illumination, complex backgrounds, and partial occlusions. Disease stages also vary in texture and color intensity, which may affect recognition accuracy. To address these, future work will incorporate image normalization, shadow removal, and adaptive thresholding to enhance robustness. Additionally, model compression and pruning will be explored to reduce computational overhead for edge deployment on mobile or drone-based systems.
The developed hybrid and explainable framework thus support practical agricultural use, enabling transparent and high-performance disease diagnostics. Future work will extend this model with real-time image capture, hyperspectral imaging, and deployment through edge computing devices or mobile applications to support precision agriculture and smart farming environments.


