Изучение функциональной значимости конфликтного варианта гена RAD51D при раке молочной железы
Автор: Молоков А.Ю., Гервас П.А., Коллантай О.В., Дударь Г.Е., Ванг Л., Хуан Ч., Чойнзонов Е.Л., Чердынцева Н.В.
Журнал: Сибирский онкологический журнал @siboncoj
Рубрика: Лабораторные и экспериментальные исследования
Статья в выпуске: 1 т.25, 2026 года.
Бесплатный доступ
Патогенные мутации в генах репарации ДНК (таких как BRCA1/2, RAD50, RAD51D, PTEN и др.) ответственны за развитие наследственного рака молочной железы и яичников. Большое количество вариантов, выявляемых с помощью технологии NGS, имеют неизвестное или конфликтное клиническое значение. Реклассификация данных вариантов играет решающую роль в рутинной лабораторной практике. Целью исследования явилась реклассификация варианта конфликтного значения гена RAD51D (rs145309168), обнаруженного у молодой пациентки бурятского происхождения с раком молочной железы, с использованием нонсенс-опосредованного распада мРНК (NMD) с последующим секвенированием по Сэнгеру. Материал и методы. Полноэкзомное секвенирование (WES) было выполнено на ДНК, выделенной из цельной крови 16 пациенток с раком молочной железы бурятского этноса, у которых отсутствовали мутации в генах BRCA1/2 (данные не представлены). Диагноз пациенток подтвержден морфологически (T1–3N0–2M0). У всех обследованных женщин был диагностирован инвазивный (протоковый) рак молочной железы неспецифического типа. Патогенные варианты, ассоциированные с заболеванием, не выявлены. Далее были проанализированы редкие варианты (MAF-0,005) для оценки их влияния на сплайсинг РНК с использованием биоинформатических инструментов, таких как SpliceAI, ESEFinder, RESCUE-ESE и EX-SKIP. Редкий миссенс-вариант в гене RAD51D (rs145309168) был идентифицирован у 39-летней пациентки бурятского этноса с раком молочной железы. Замороженные лейкоциты этой пациентки были разделены на две группы: экспериментальную и контрольную. Образцы культивировали в течение 5–6 дней и обрабатывали пуромицином (только экспериментальную группу) в течение 4–6 ч перед выделением РНК для предотвращения NMD с последующим секвенированием по Сэнгеру. Результаты. In vitro эксперименты проводились на живых лейкоцитах пациентки с раком молочной железы, имеющей вариант c.932T>A гена RAD51D. Ампликоны кДНК были получены из РНК, выделенной из контрольных и экспериментальных лейкоцитов (обработанных пуромицином для предотвращения деградации, опосредованной NMD). Для точной оценки аберраций сплайсинга транскрипты экспериментальных лейкоцитов сравнивались с транскриптами контрольных культур лейкоцитов с помощью секвенирования по Сэнгеру. Последовательности транскриптов кДНК сравниваемых образцов в обоих случаях сохраняют изучаемый вариант, что указывает на то, что вариант не активирует NMD и, следовательно, не влияет на сплайсинг. Заключение. В данном исследовании впервые представлен in vitro анализ варианта RAD51D (rs145309168), найденного у молодой пациентки бурятского этноса с раком молочной железы. Наши экспериментальные данные демонстрируют, что вариант c.932T>A не нарушает нормальный сплайсинг, что служит основанием для реклассификации данного «Конфликтного варианта» на «Вероятно доброкачественный», что согласуется с литературными данными и данными ранних классификаций.
МРНК, ген репарации ДНК, RAD51D, рак молочной железы, этнические группы, буряты, неевропеоиды
Короткий адрес: https://sciup.org/140314346
IDR: 140314346 | УДК: 618.19-006.6:575.113 | DOI: 10.21294/1814-4861-2026-25-1-46-53
Functional analysis of the RAD51D gene conficting variant in breast cancer
Germline pathogenic variants in DNA repair genes (BRCA1/2, RAD50, RAD51D, PTEN and etc.) are responsible for the development of hereditary breast and ovarian cancers. The large number of variants detected by NGS technology have unknown or conficting clinical signifcance. Reclassifcation of these variants plays a crucial role in their application in routine laboratory practice. The aim of the current study was to reclassify conficting RAD51D gene variant (rs145309168) in a young Buryat breast cancer patient using the translation-dependent nonsense-mediated mRNA decay (NMD) pathway. Material and Methods. Wholeexome sequencing (WES) was performed in the germline DNA of 16 non-BRCA BC Buryat patients (data not shown). The diagnosis in all patients was confrmed morphologically (T1–3N0–2M0). All tested women were diagnosed with invasive (ductal) carcinoma of no special type. Rare variants (MAF-0.005) were analyzed to assess their impact on the RNA splicing using in silico tools like SpliceAI, ESEFinder, RESCUE-ESE, and EX-SKIP. A rare missense variant in the RAD51D gene (rs145309168) was identifed in a 39-year-old Buryat breast cancer patient. Frozen patient leukocytes were divided into experimental and control samples. The samples were cultured for 5–6 days and treated with puromycin (only experimental samples) for 4–6 hours prior to RNA isolation to avoid NMD followed by Sanger sequencing. Results. In vitro experiments on live leukocytes from a breast cancer patient with the c.932T>A variant of the RAD51D gene were conducted. cDNA amplicons were obtained from RNA isolated from control and experimental leukocytes (treated with puromycin to avoid nonsense-mediated decay). For an accurate assessment of splicing aberrations, transcripts from the experimental leukocytes were compared to transcripts from control leukocytes by using Sanger sequencing. In both cases, the presence of the studied variant in the RNA signifed that the variant did not activate NMD and therefore did not affect splicing. Conclusion. This study presents the frst in vitro functional analysis of the RAD51D variant (rs145309168) identifed in a young Buryat breast cancer patient. Our experimental data demonstrate that this variant does not disrupt normal splicing, providing evidence for its reclassifcation as «Likely Benign», which is consistent with published data and previous classifcations.
Текст научной статьи Изучение функциональной значимости конфликтного варианта гена RAD51D при раке молочной железы
Breast cancer (BC) is the most common cancer among women globally. Approximately 5–10 % of all breast cancers are hereditary [1]. Germline pathogenic variants in DNA repair genes, notably BRCA1/2, RAD51C/D, PTEN, and others, significantly increase lifetime breast and ovarian cancer risk. The presence of the pathogenic variant offers certain advantages in treatment (personalized treatment using PARP inhibitors), enables disease prognosis, and helps assess the risk of BC among family relatives.
However, only 2–15 % of patients tested for mutations carry pathogenic variants, with the remainder having variants of uncertain significance (VUS) or table/Таблица
sanger sequencing PCR primers for conflicting variant of gene RAD51D rs145309168 (c.932t>a:p.ile311asn)
Праймеры для секвенирования по Сэнгеру конфликтного варианта гена RAD51D rs145309168 (c.932t>a:p.ile311asn)
PCR primers/ПЦР праймеры
Nucleotide sequence/ Нуклеотидная последовательность
RAD51D-cDNA-F
5'- CAAACCTGCCCTCGGAC-3'
RAD51D-cDNA-R
5'- TTTCTGGGTCCTCGCAAT-3'
Примечание: таблица составлена авторами.
Note: created by the authors.
conflicting pathogenicity classifications. VUS may pose a significant diagnostic challenge. On the other hand, VUS in genetic testing may be upgraded to Likely Pathogenic or Pathogenic as more evidence becomes available through research (familial segregation analysis, re-evaluation) [2]. The primary research method for VUS classification currently involves analyzing the variant frequency in population, which is not applicable to small understudied ethnic groups or rare variants. Therefore, the alternative approaches are necessary to reclassify variants ( in silico or in vitro analysis). In recent years, there has been a steady increase in the number of bioinformatics tools ( in silico ) for predicting variant pathogenicity. Despite advances in the development of in silico algorithms for predicting variant effects, functional analysis modeling the effect at the transcriptional or post-transcriptional level remains the classical method to confirm the pathogenicity of variants. One such system is the translation-dependent nonsense-mediated mRNA decay (NMD) pathway that recognizes and eliminates mRNAs encoding truncated and full-length proteins with no or undesired functions [3].
In our study, we focused on the conflicting RAD51D gene variant (rs145309168). Our previous research has shown that mutations in the RAD51D gene play a significant role in the pathogenesis of breast cancer in Buryat [4, 5]. Furthermore, data from the literature indicate that RAD51D gene variant (rs145309168) is rare (allele frequency is 0.003298 in Asian populations compared to 0.00007798 in Caucasian populations) and its pathogenetic significance remains conflicting due to a lack of in vitro studies [6–10].
The aim of the current study was to reclassify conflicting RAD51D gene variant (rs145309168) in a young Buryat BC patient using the translationdependent nonsense-mediated mRNA decay (NMD) pathway followed by Sanger sequencing.
Material and Methods
The study included 16 Buryat BC patients (the median age was 37 ± 7.94 years) who were treated at the Cancer Research Institute of the Tomsk National Research Medical Center. The diagnosis in all patients was histologically confirmed (T1–3N0–2M0). All tested women were diagnosed with invasive (ductal)
carcinoma of no special type. All patients provided signed informed consent to participate in the study.
Whole‐exome sequencing (WES) was performed in the germline DNA of non- BRCA patients (data not shown). Rare variants (MAF<0.005) were analyzed using in silico tools to assess their impact on the RNA splicing. Disruption/creation of splice sites was evaluated using SpliceAI, ESEFinder, and RESCUE-ESE. The EX-SKIP were used to examine the effects on exonic splicing enhancer (ESE) and silencer binding sites (ESS) that play a crucial role in correct splice-site identification or relevant regulation of splicing [11].
Frozen patient leukocytes were divided into experimental and control samples. The samples were cultured for 5–6 days and treated with puromycin (only experimental samples) for 4–6 hours prior to RNA isolation to avoid nonsense-mediated decay (NMD) [12]. Total RNA was isolated using the RNAeasy kit (Qiagen) according to the manufacturer instructions. A 1.0 µg of total RNA was reverse transcribed using random primers and SuperScript II reverse transcriptase (Invitrogen, Carlsbad, CA, USA). For an accurate assessment of splicing aberrations, transcripts from the experimental leukocytes were compared to transcripts from control leukocyte cultures using Sanger sequencing (Figure 1). PCR primers were designed according to the ENIGMA recommendations (Table).
Results
A rare missense variant in the RAD51D gene was identified in a 39-year-old Buryat patient diagnosed with stage T2N3M0 left-sided invasive luminal B breast carcinoma using whole-exome sequencing analysis. The identified variant (c.932T>A, p.Ile311Asn, rs145309168) of the RAD51D gene resides on chromosome 17 (chr 17:35101008A>T (hg38)) and leads to an amino acid substitution at the protein level. According to the gnomAD database (v4.1.0), the minor allele frequency is 0.0001 [6]. However, in the dbPubMed ClinVar database, this substitution has been classified as a variant of conflicting significance (reported from a variant of uncertain to benign significance). In silico analysis provides that it can be likely deleterious (PrimateAI-3D is 0.77), or moderate pathogenic (SIFT). According to the ESEfinder and RESCUE-ESE tools, the studied RAD51D gene variant affects exonic
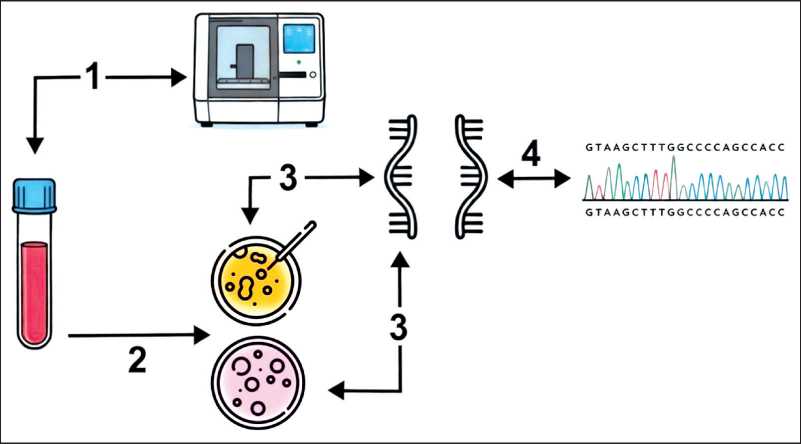
Fig. 1. Design of the study. 1. Buryat female patients (37 ± 7,94 years) with BC were tested using NGS; 2. The patient samples were cultured for 5–6 days and treated with puromycin (only experimental samples) for 4–6 hours before RNA extraction to inhibit NMD; 3. Total RNA was isolated using the RNeasy Kit (Qiagen). Reverse transcription was performed with random primers and SuperScript II Reverse Transcriptase (Invitrogen, USA); 4. Transcripts from the experimental leukocytes were compared to transcripts from control leukocyte cultures using Sanger sequencing. Note: created by the authors
Рис. 1. Дизайн исследования. 1. Материал пациенток бурятской национальности (37 ± 7,94 года) с диагнозом рак молочной железы изучен с использованием технологии NGS. 2. Образцы лейкоцитов пациентки культивировали в течение 5–6 дней и обрабатывали пуромицином (только экспериментальные образцы) в течение 4–6 ч перед экстракцией РНК для ингибирования NMD. 3. Тотальную РНК выделяли с помощью набора RNeasy Kit (Qiagen). Обратную транскрипцию проводили с использованием случайных праймеров и обратной транскриптазы SuperScript II (Invitrogen, США). 4. Транскрипты кДНК из экспериментальных лейкоцитов сравнивали с транскриптами кДНК из контрольных культур лейкоцитов пациентки с помощью секвенирования по Сэнгеру. Примечание: рисунок выполнен авторами
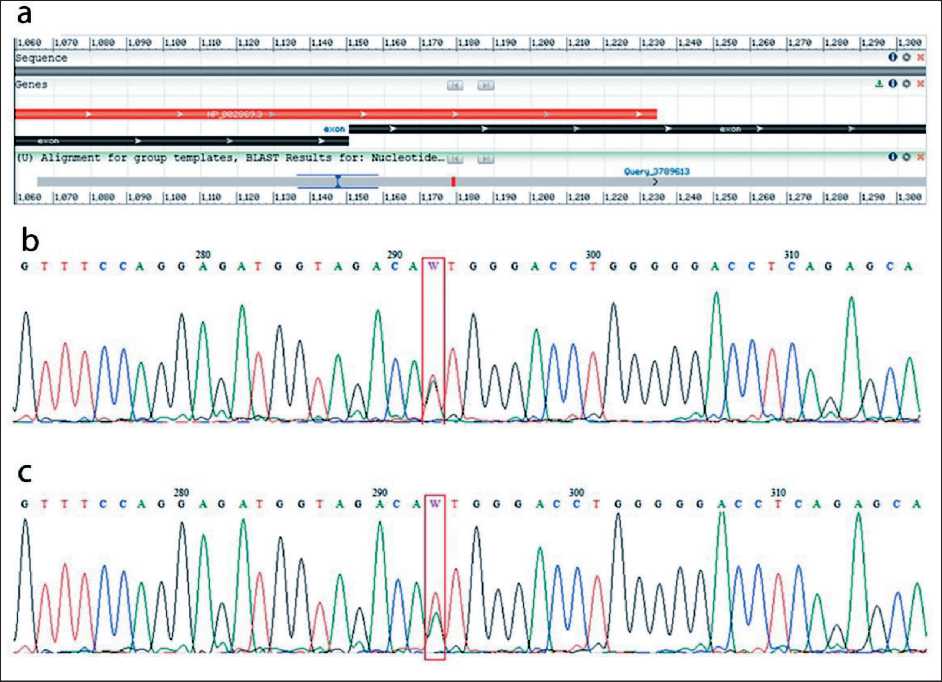
Fig. 2. Sequencing chromatogram of RAD51D gene cDNA (rs145309168), in vitro functional assays. а – cDNA sequencing chromatogram, obtained via Nucleotide BLAST, shows an inter-exonic region, the mutation site (red); b – cDNA sequencing chromatogram of the sample without puromycin treatment shows the c.932T>A (W) substitution in the RAD51D gene; c – cDNA sequencing chromatogram of the puro-mycin-treated sample (with NMD inhibited) also shows the c.932T>A (W) substitution in the RAD51D gene. Note: created by the authors Рис. 2. Хроматограмма секвенирования кДНК гена RAD51D (rs145309168), функциональные исследования in vitro : а – хроматограмма секвенирования кДНК, полученная с помощью Nucleotide BLAST, показывающая межэкзонный регион и место мутации (красный); b – хроматограмма секвенирования кДНК контрольного образца, без обработки пуромицином, показывающая замену c.932T>A (W) в гене RAD51D ; c – хроматограмма секвенирования кДНК экспериментального образца, обработанного пуромици-ном (с ингибированием NMD), также показывающая замену c.932T>A (W) в гене RAD51D .
Примечание: рисунок выполнен авторами splicing enhancers (ESE) playing a crucial role in correct splice-site identification or relevant regulation of splicing (ESEfinder tools/ESE2/, edu/cgi-bin/tools/ESE3/ and RESCUE-ESE . We performed a functional analysis of the c.932T>A mutation in the RAD51D gene, which plays a key role in homologous recombination.
In our study, we conducted in vitro experiments on live leukocytes from a breast cancer patient with the c.932T>A variant of the RAD51D gene to assess nonsense-mediated RNA degradation. cDNA amplicons were obtained from RNA isolated from control and experimental leukocytes. The primers were designed for the inter-exon region between exons 9 and 10, as shown in Figure 2a. For an accurate assessment of splicing aberrations, transcripts from the experimental leukocytes (treated with puromycin to avoid nonsense-mediated decay) were compared to transcripts from control leukocyte cultures by using Sanger sequencing (Figure 2b-c). Figure 2b-c shows the transcript sequences of the compared samples (leukocytes treated (experimental) and untreated (control) with purimycin). In both cases, the RNA retains the presence of the studied variant, indicating that the variant does not activate NMD and does not affect splicing. If the variant affects splicing (pathogenic), nonsense-mediated degradation destroys the amplicon containing the damaging variant.
We found that this variant did not affect splicing and did not activate the nonsense-mediated RNA degradation mechanism.
Discussion
Our study was dedicated to investigating hereditary breast cancer in the Buryat ethnic group, as the absence of BRCA1/2 gene mutations had previously been demonstrated in this population. Furthermore, we had earlier identified the pathogenic variant c.757C>T in the RAD51D gene, which was subsequently classified as a founder mutation for the Buryat ethnic group [3, 4]. According to Yao et al., c.757C>T in the RAD51D gene is a founder variant in the Chinese population, and its presence serves as an indication for the successful use of PARP inhibitors [13]. Therefore, other alterations in the RAD51D gene may significantly contribute to the pathogenesis of hereditary breast cancer in the Buryat ethnic group. In this study, we continued the search


