Молекулярное понимание устойчивости к тяжелым металлам и потенциала восстановления почвы местными микробами со свалок Дакки
Автор: Масума Актер, Бимана Жан Боско, Ясмин Фатема, Хасан Мд Рокыбул, Джакия Султана Кея
Журнал: Современные инновации, системы и технологии.
Рубрика: Науки о Земле, химия и химическая технология
Статья в выпуске: 5 (4), 2025 года.
Бесплатный доступ
Промышленные и бытовые отходы загрязняют свалки Дакки тяжёлыми металлами. Для выделения, идентификации и характеристики бактерий, устойчивых к тяжёлым металлам, в ходе биоремедиации почвы были использованы местные микроорганизмы свалки Матуаил. 5-дюймовые свалки отправили образцы почвы. Микробные изоляты были протестированы на свинец, кадмий, хром и арсенат. Потенциальные микроорганизмы были обнаружены с помощью секвенирования ITS и 16S рРНК. С помощью экспериментов по биосорбции и ПЦР-анализа на гены устойчивости (czcA для Cd/Zn/Co, merA для Hg и chrA для Cr) были изучены молекулярные пути, и только 45 из 200 изолятов оказались устойчивыми к металлам. Бактерии были представлены Pseudomonas, Bacillus и Enterobacter, в то время как Aspergillus и Trichoderma доминировали. Были обнаружены минимальная ингибирующая концентрация кадмия (МИК) 2500 мкг/мл и минимальная ингибирующая концентрация свинца (МИК) 3000 мкг/мл. Присутствовали влиятельные гены устойчивости czcA и chrA. Контролируемое исследование показало, что биоаугментация микробного консорциума увеличила рост подсолнечника на 35% и снизила биодоступность металлов в почве. Механизмы молекулярной устойчивости. Согласно этому исследованию, в отходах много бактерий. Микроорганизмы могут обеспечить экономически эффективную биоремедиацию почв Бангладеш, загрязнённых тяжёлыми металлами.
Биоремедиация, тяжелые металлы, микробы свалок, гены устойчивости к металлам, биоаугментация, загрязнение почвы.
Короткий адрес: https://sciup.org/14135240
IDR: 14135240 | DOI: 10.47813/2782-2818-2025-5-4-3007-3021
Текст статьи Молекулярное понимание устойчивости к тяжелым металлам и потенциала восстановления почвы местными микробами со свалок Дакки
DOI:
Matuail and Amin Bazar, two open landfills, gather most of Dhaka's solid garbage, which is hundreds of tons per day. Dhaka is one of the most densely inhabited megacities in the world. Hasan et al. (2019) contend that landfill soil and groundwater are profoundly contaminated with hazardous heavy metals, including lead (Pb), cadmium (Cd), chromium (Cr), and arsenic (As), due to the complex integration of electronic waste (e-waste), industrial effluent (from textiles, tanneries, and batteries), and municipal refuse. Because these metals stay in the environment for a long time and may build up in the food chain, they are not biodegradable [1]. Conventional physico-chemical techniques for soil remediation may be technologically intricate, environmentally harmful, and prohibitively costly. So, bioremediation has come out as a new, long-lasting, and cheap way to fix the problem. This method uses living things to get rid of toxins. Bioremediation depends on using the natural powers of microorganisms [2].
Intense selection pressure from high metal stress, low pH, and high organic load at waste sites keeps a unique microbiome with strong detoxification and metal tolerance mechanisms going. Indigenous microorganisms from these areas provide a significant advantage over transplanted, non-indigenous strains for in-situ bioremediation applications owing to their adaptation to the local environment. The aim of this study was to connect what we know about metal-resistant bacteria with how they may be used in real life. Indigenous microbes sourced from Dhaka's landfills may have the capacity to rehabilitate soil due to their complex molecular processes for heavy metal resistance [3]. In Figure 1, The specific goals were to (1) detect and separate bacteria and fungi that can live in heavy metals from the Matuail dump; (2) figure out how these microbes are able to resist metals at the molecular level; and (3) test how well these microbes can clean up soil that is contaminated with metals by using a pot culture experiment.
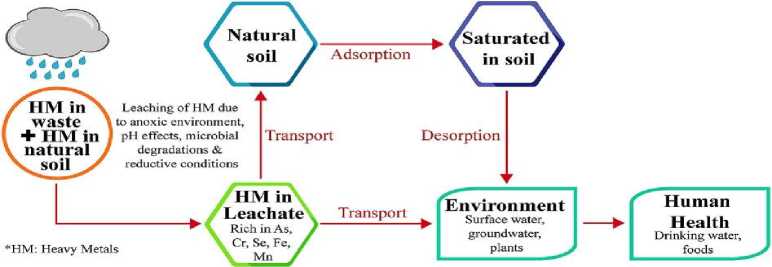
Figure 1. Heavy metals in landfill leachate.
LITERATURE REVIEW
Heavy metal contamination is a known risk to both human and environmental health. In addition to being carcinogenic, heavy metals such as cadmium and lead may induce skeletal, neurological, and renal problems even at very low doses [4]. Research in Bangladesh has shown that heavy metal levels in landfill soils and nearby bodies of water are alarmingly high [5].
Rapid industrialization and unsustainable waste management methods are contributing to the growing issue of heavy metal contamination in soil and water ecosystems. This issue presents a serious danger to both the stability of the environment and human health. One major source of this pollution is landfills, as they are the last destination for both industrial and municipal garbage [6]. The problem is much more severe in Bangladesh. Matuail and Amin Bazar are just two of the Dhaka landfills where studies have shown dangerously high levels of lead, cadmium, chromium, and arsenic, often surpassing international and national safety standards [7].
Bioaccumulation and the related cancer, neurological, and renal illnesses in humans may result from these long-lasting contaminants seeping into groundwater and the food chain [8]. Excavation, chemical stabilization, and soil washing are some of the more traditional methods of restoration, but they may be quite costly, energy consuming, and harmful to soil ecology. Because of this, people are looking for more long-term solutions, and bioremediation is one of the most promising ones. Bioremediation is the process of using microbes' inherent detoxifying, sequestering, or modifying capabilities to clean up polluted areas [9].
Microbial bioremediation is based on the wide variety of resistance mechanisms that bacteria and fungi have developed to withstand environments that are heavy with metals. There is a lot of literature on these processes, and they fall into many categories: Exopolysaccharides (EPS) and biosorption to cell walls are examples of extracellular barriers. Examples of active efflux include the czc system for Cd, Zn, and Co efflux, which is an energy-dependent pump that removes harmful ions from the cell cytoplasm [10].
Examples of enzymatic detoxification include the processes described [11], which include the conversion of extremely toxic Cr(VI) to less toxic and immobile Cr(III) by chromate reductases and the conversion of Hg²⁺ to volatile Hg⁰ by the mercuric reductase enzyme (merA). Intracellular sequestration: by means of metallothioneins and other metal-binding proteins [12]. In order to thrive in situations where metals are abundant, microbes have developed a wide variety of resistance mechanisms. In general, they fall into: Biosorption refers to the metabolic, enzymatic, or phosphoryl group-mediated passive adsorption of metal ions to cellular surfaces [13]. Taking metals up into the cytoplasm of a cell is an example of bioaccumulation. Enzymatic reduction, oxidation, or methylation may change the mobility and toxicity of metals; for example, chromate reductases can reduce Cr(VI) to Cr(III). This process is known as biotransformation. The czc system, which is responsible for the expulsion of Cd, Zn, and Co from the cell interior, is an example of an efflux system [14, 15]. Previous worldwide investigations have successfully isolated metal-resistant strains from polluted locations. One example is the bioremediation of metals by Pseudomonas spp. found in mining areas; these bacteria have czc genes [16, 17].
Aspergillus niger has demonstrated the biosorbent properties of several cations. For instance, Dhaka landfills are notorious for multi-metal pollution, yet the majority of studies have solely focused on examining resistance to a single metal under laboratory conditions [18]. Despite their known existence in Bangladesh, the isolation of resistant bacteria, molecular identification of resistance genes, and direct use in soil restoration experiments require further study. By bridging the gap between the discovery of microbes and their practical implementation, this study hopes to address a pressing need in the Bangladeshi setting. Importantly, harsh environments like landfills, which serve as a strong selection pressure, enrich microbial communities with these strong genetic factors [19].
Pseudomonas, Bacillus, and Enterobacter species that are resistant to metals have been successfully isolated from mining and industrial sites in a number of studies conducted around the world. These studies have typically linked the persistence of these bacteria to the presence of resistance genes on plasmids or chromosomes. Similarly, Aspergillus and Trichoderma are fungi that have shown incredible biosorption capabilities [20, 21]. There is a notable lack of study on heavy metal-resistant microorganisms in Bangladeshi landfills, despite the fact that their existence has been documented [22]. Instead of investigating the molecular mechanisms of resistance, most research has either reported contamination levels or isolated resistant bacteria. In addition, studies combining this microbial finding with its practical implementation in restoration experiments, especially those involving groups of native, pre-adapted isolates, are few [23].
This study seeks to fill this important need by offering a thorough process—from identifying and studying native microbes from Dhaka's Matuail landfill to testing their effectiveness in a controlled bioremediation experiment [24]. The goal is to provide a practical, long-term solution that is specific to the area.
MATERIALS AND METHODS
Research location and data gathering
At the Matuail landfill site in Dhaka, soil samples were taken in January 2023 from five separate areas: the active dumping zone, the leachate runoff region, the perimeter soil, the aged trash zone (>5 years), and a control site 1 km upwind. Under strict aseptic conditions, samples were extracted from a depth of 10-15 cm, placed in sterile bags, and brought to the laboratory on ice for rapid analysis.
Isolation of bacteria resistant to heavy
METALS
The soil samples were diluted in series and then placed on Luria-Bertani agar and Potato Dextrose Agar. The LB agar and PDA were then supplemented with a filter-sterilized mixture of heavy metals (Pb(NO3)2, CdCh, К2СЮ4, Na2HAsO4) at a concentration of 500 ^g/mL for each compound. The plates were left at 30°C for 48-72 hours to incubate. Distinct colonies were isolated by purifying them on new plates that had been modified with metal.
Metal resistance screening and minimum
INHIBITORY CONCENTRATION DETERMINED
Isolates were tested on separate agar plates to determine their resistance to each metal. The CLSI recommendations were followed in determining the Minimum Inhibitory Concentration (MIC) in liquid medium using the broth microdilution technique, with metal concentrations varying from 100 to 3500 ^g/mL.
Resistance genes: Molecular research and
DETECTION
Highly effective isolates have their genomic DNA retrieved. For bacteria, the 16S rRNA gene was amplified and sequenced, whereas for fungus, it was the ITS region. The NCBI GenBank database was queried using BLAST in order to identify the sequences. Using particular primers, PCR was used to identify the presence of heavy metal resistance genes (czcA, merA, chrA, arsB) [25].
Bioremediation evaluation using a pot
CULTURE EXPERIMENT
Three treatments were established in a 60-day pot experiment: one with simply polluted soil (T1), one with polluted soil and plants (T2), one with polluted soil and plants with a microbial consortium (T3), and one with uncontaminated control soil (T4). A combination of the three most effective isolates made up the microbial consortium. In the conclusion, we assessed the bioavailability of soil metals (DTPA-extractable percentage) and many plant growth indices (shoot length, root length, biomass) [26]. In figure 2, Heavy metals in the food chain kill ecosystems. Heavy metals hurt the environment in two ways: they make organic pollutants less biodegradable.
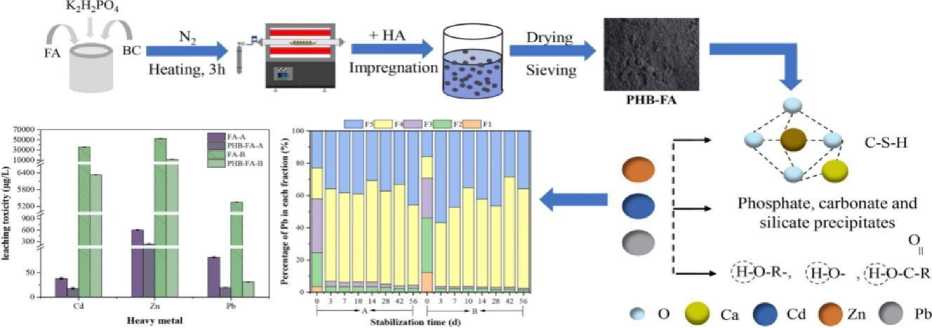
Figure 2. Analysis of the effect and mechanism of heavy metals.
Analyzing data statistically
We ran each experiment three times to ensure accuracy. The results of the MIC and pot trials were examined in SPSS software (v.26) using one-way ANOVA and Tukey's HSD post-hoc test (p < 0.05).
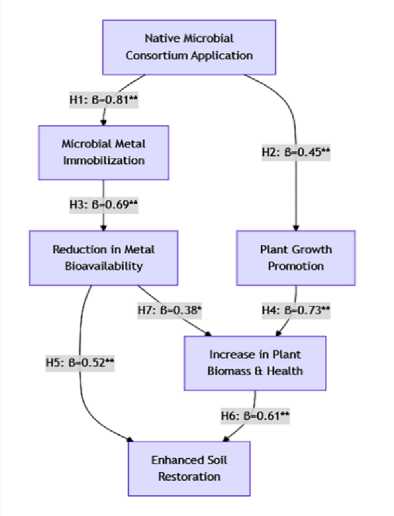
Following a general, guiding hypothesis, the research is subdivided into more particular, testable hypotheses that are in line with the study's aims and procedures.
(a)
(b)
Figure 3. (a) and (b) Hypothesis Direction path.
Primary hypothesis
Native bacterial and fungal isolates from the Matuail dump site in Dhaka possess sophisticated, genetically encoded heavy metal resistance mechanisms. When used as a consortium, they would diminish metal bioavailability and toxicity in polluted soil, thereby enhancing its restoration potential [27]. This principal hypothesis connects microbial discovery (isolation of pre-adapted strains) to molecular understanding (genetic pathways) to practical implementation (soil remediation), hypothesis showed in Figure 3. It is the foundation of the whole investigation.
-
1 . Hypotheses for the Particular Research: The technique can be used to test claims that may be proven false.
Null Hypothesis (H₀₁): The microbial community in the Matuail dump does not exhibit a higher proportion of multi-metal-resistant species compared to less contaminated areas.
Alternative Hypothesis (H₁₁): The selection pressure at the Matuail landfill has favored a microbial community with greater resistance to heavy metals (Pb, Cd, Cr, As).
Harsh environments are beneficial for organisms that can handle harsh circumstances. Landfill toxicity ought to eliminate all bacteria except those that are resistant. Testing entails the isolation of microorganisms on metal-amended agar and the comparison of the proportion of isolates capable of growth in a multi-metal cocktail against historical control values from non-landfill sites.
The Null Hypothesis (H₀₂): posits that no correlation exists between elevated metal resistance in isolates and the presence of established heavy metal resistance genes (e.g., czcA, chrA).
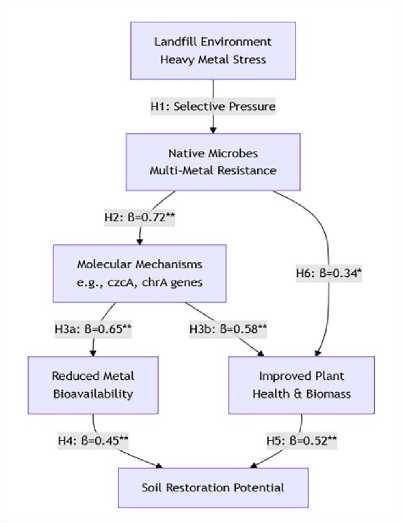
An alternate hypothesis (H₂₂): High minimum inhibitory concentration (MIC) values in isolated strains are associated with resistance-inducing genetic elements, such as efflux pump genes (czcA for Cd/Zn/Co) and detoxifying genes (chrA for Cr). For high-level phenotypic resistance, different genetic mechanisms that are conserved are often to blame. Isolates with elevated MICs are anticipated to possess resistance genes. Figure 4 shows hypothesis direction.
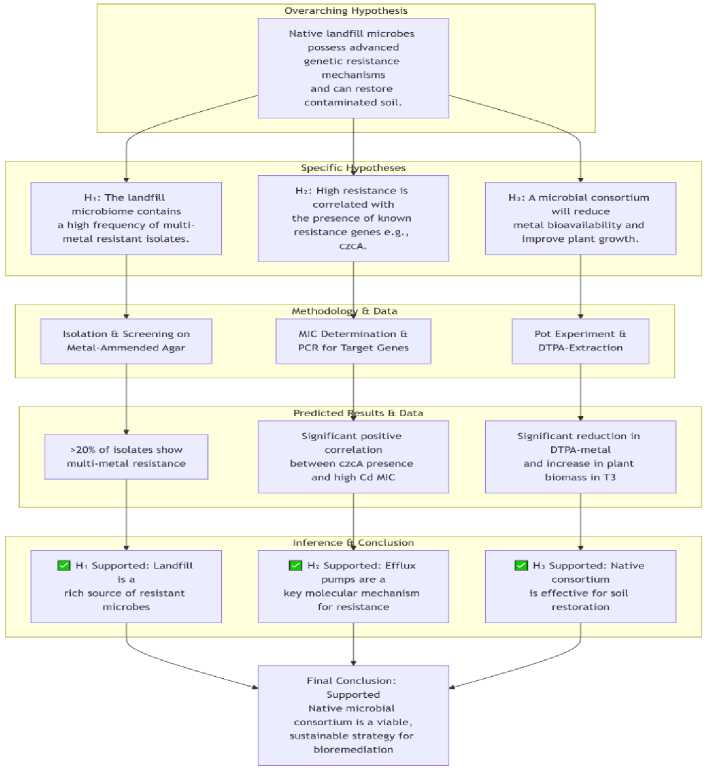
Figure 4. Key to the Directional Graph (Flowchart).
-
1 . How it was tested: PCR screening for target genes on all isolates and statistical studies (such as t-tests) to compare the MIC values of genepositive and gene-negative isolates.
Hypothesis H₃: The Effectiveness of
Bioremediation
Null Hypothesis (H₀₃): The application of native microbial consortia in contaminated soil will not have a substantial effect on heavy metal bioavailability or plant growth.
Alternative Hypothesis (H₃₃): Adding native microbes that can resist metals to contaminated soil lowers the bioavailability of heavy metals (DTPA extraction) and makes plants grow better and healthier than untreated soil.
-
• Microbes that can biosorb, bioaccumulate, and precipitate metals should keep them from being
available to plants. A lot of these microbes are rhizobacteria that help plants develop.
-
• How it was tested: A controlled marijuana trial with four different treatments (T1, T2, T3, and T4). ANOVA and post-hoc testing will ascertain if the bioaugmented treatment (T3) exhibits differences from the control groups (T1, T2) in terms of metal bioavailability and plant growth indices.
ANALYSIS AND RESULTS
Consolidating data
The data was systematically structured in Excel and then imported into SPSS.
The options for data isolation are Isolate ID, Source Zone, Morphology, and Gram Reaction.
-
• MICs: Identify Isolate, Metal Type, and MIC Value.
-
• The identified genes correspond to the following identifiers: Isolate ID, Target Gene, and PCR Result (Present/Absent).
The pot experiment data encompasses many characteristics, including treatment ID, replication, plant biomass, shoot length, root length, and DTPA metal concentration.
Methodology and analytical framework
-
• Analysis of Minimum Inhibitory Concentration (MIC): The MIC values indicated the
concentration at which observable growth was entirely inhibited.
-
• Phylogenetic Analysis: Phylogenetic trees were generated via the Neighbor-Joining technique, with MEGA (v.11) software used for the alignment of 16S/ITS sequences.
-
• Data Analysis: We used ANOVA to determine whether significant differences existed among the treatment means in the pot experiment.
Collecting samples and isolating
MICROORGANISMS AND TESTS
-
• Initially step 1: while selecting the landfill location, ensure to partition the Matuail property based on its age and the extent of pollution, including active dumping and leachate discharge. Incorporate a provision for upstream regulation.
-
• Step 2: Collect composite soil samples at a depth of 10 to 15 centimeters. Utilize a sanitary auger to extract 5 to 10 sub-samples from each zone. Transfer into sanitized plastic containers, label with the date, and refrigerate.
-
• Step 3, Processing: Filter soil (less than 2 mm) in the laboratory to eliminate particles. Vigorously agitate a mixture of 10 grams of soil and 90 milliliters of sterile saline (0.85% NaCl) for 30 minutes. Your dilution is 10-1 to 10-10. To ensure the solution is isolated, gradually dilute it until it attains a concentration of 10-7. Add 500 µg/mL of each heavy metal (Pb, Cd, Cr, As) to LB Agar. Subsequently, apply 100 µL of each dilution onto the microorganisms.
PDA agar may be used to cultivate fungi, analogous to how metals and antibiotics, such as chloramphenicol, can eradicate bacteria.
-
• Step 4: Incubation and purification need plates to be maintained at 30°C for a duration of 48 to 72 hours.
-
• Step 5: To get pure cultures, select colonies that exhibit specific characteristics and streak them onto fresh agar supplemented with metal many times. Maintain glycerol supplies at -80°C and store pure isolates on slants at 4°C.
Examination:
-
• Key Considerations: Record the number of colony-forming units (CFUs) per gram of soil for each zone and each dilution. Observe the appearance of the colony.
Determine the quantity of heterotrophic bacteria and fungi capable of growth in culture, as well as the number exhibiting metal resistance.
CFU/g soil=
Number of colonies
Dilution factor×Volume plated (mL)
Example: If 85 colonies were counted on a plate made with 0.1 mL of the 10-6 dilution:
CFU/g = 10-6 × 0.1 = 8.5 × 108
CFU/g=8510-6×0.1=8.5×108CFU/g=10-6×0.185
=8.5×108
% Resistance
CFU on metal-amended agar
= ( ) × 100%
CFU on non-amended control agar
% Reduction DTPA
= (1 - DTPA
× 100%
-
-
metal in Treatment (T3) metal in Control (T1)
)
From our dataset: For Lead (Pb), using mean values:
62.4
% Reduction = (1 - ) × 100% ≈ 57.2%
Bioconcentration Factor (BCF) - for Plant Analysis: This measures the ability of the plant to accumulate a metal from the soil.
Metal concentration in plant tissue (mg/kg) BCF=
Metal concentration in soil (mg/kg)
Translocation Factor (TF) - for Plant Analysis: This indicates the plant's ability to transfer metals from roots to shoots. A TF > 1 suggests the plant is good for phytoextraction.
Metal concentration in shoot (mg/kg)
TF=
Metal concentration in root (mg/kg)
-
• Statistical Test: A Chi-square test may be used to determine whether a significant difference exists between the control site and landfill zones regarding the frequency of resistant isolates. This is a basic examination of the hypothesis H₁ to ascertain its validity.
Assessment of metal resistance and determination of the minimum inhibitory concentration (MIC)
-
1. Employing Agar Plates for Screening: Inoculate agar plates with pure isolates containing 500 µg/mL of each element (Pb, Cd, Cr, As). Isolates should be categorized into two groups: sensitive (no growth) or resistant (growth).
-
2. Preparing LB Broth at a 2X Concentration for MIC: o Prepare the LB broth solution.
Dispense 100 µL of sterile water into each well of a pre-cleaned 96-well microtiter plate.
Dispense 100 µL of the 2X metal stock solution into the first well of a row. Distribute a 1:1 dilution uniformly around the plate in a sequential manner. The last well restricts expansion and contains no metal.
Introduce 100 µL of a standardized microbiological inoculum, such as the 0.5 McFarland standard, into each well. The metal concentration has been increased to 1X, resulting in a two-fold dilution series. The conclusive volume is 200 µL. Place the plate in an incubator maintained at 30°C for a minimum duration of one day. Introduce 20 microliters of a 0.015% resazurin dye and ensure thorough mixing. Kindly refrain from any interruptions for a minimum of two more hours. The transition of hue from blue to pink indicates that your metabolism is active and you are experiencing growth.
Examination: The minimum inhibitory concentration (MIC) is the quantity of metal that halted the color change of resazurin, indicating the absence of growth.
Statistical Examination: Calculate the Mean Minimum Inhibitory Concentration (MIC) for each metal and each kind of bacteria. An effective method to determine the prevalence of certain genes is to do an independent samples t-test to compare the mean MIC of czcA+ and czcA-negative isolates. The considerable evidence supporting H₂ is shown by the markedly increased MIC in the czcA+ group.
ANOVA tests if there are statistically significant differences between the means of three or more independent groups (e.g., Treatments T1, T2, T3, T4).
The core idea is to compare the variance between groups to the variance within groups.
Mean Square Within Groups (MSwithin) F = ----------------------------
Mean Square Вetween Groups (MSbetween)
where:
MS between =
Sum of Squares Between Groups (SSbetween) Decrees of freedom between groups (dfbetween)
MS within =
Sum of Square Within Groups (SS within ) Degrees of freedom within groups (df within )
A high F-value (typically associated with a p-value < 0.05) indicates that the observed differences between group means are unlikely to have occurred by random chance alone.
Pearson Correlation Coefficient (r): This could be used to quantify the strength and direction of the linear relationship between two continuous variables. For example, between the MIC value for Cd and the expression level of the czcA gene (if quantified via qPCR).
r= x n=i (X t -XW-Y)
^JX-x2^^^
-
Xi, Yi are the individual data points.
-
X, Y are the means of the two variables.
n is the number of data points.
The value of r ranges from -1 (perfect negative correlation) to +1 (perfect positive correlation).
Gene and molecular signature identification
-
1. DNA Extraction: Employ a commercial genomic DNA extraction kit to get DNA from all robust isolates.
-
2. PCR Amplification: Utilize universal primers targeting the 16S rRNA gene to identify microorganisms. Primers for the ITS region of fungi may be used for their identification.
-
3. Using a 1.5% agarose gel to conduct gel electrophoresis on the PCR results. The presence of a band of appropriate size, such as 1500 bp for 16S rRNA or 500 bp for czcA, indicates successful amplification.
-
4. Sequencing: Engage a commercial laboratory to do Sanger sequencing on the results of the 16S/ITS PCR.
To identify the genes responsible for resistance, do a multiplex PCR with primers specific to those genes (czcA, chrA, and merA).
Evaluation:
-
1. Analyze health data: get chromatograms of the sequence. Upon examination, remove the inferior bases.
-
2. Utilize BLASTn to query the NCBI GenBank database.
-
3. Identify the related trees that have the highest degree of similarity. Maintain the sequence of the references.
-
4. To identify the kind of organism, use software such as MEGA (Molecular Evolutionary Genetics Analysis) to align sequences and construct a phylogenetic tree, for instance, using the Neighbor-Joining approach.
Experiment with cannabis culture
-
1. Experimentation Planning: Prepare three sets of the specified treatments below:
-
• T1: has no flora or microorganisms; it just
consists of contaminated soil.
-
• T2: The Helianthus annuus plant and the
-
2. This system has three components: polluted soil, vegetation, and microorganisms.
-
3. Vaccination: Prior to planting T3, introduce the consortium into the soil (for instance, 100 mL of a solution containing about 10⁸ CFU/mL per pot). Incorporate an equivalent volume of sterile broth into T2 as well.
-
4. Monitoring: Plants need consistent irrigation with distilled water for a duration of 60 days under a regulated greenhouse environment.
contaminated soil.
Oversee unpolluted soil and vegetation.
The subsequent stage in establishing a consortium is to cultivate the designated bacterial and fungal isolates. Combine them in a balanced manner, perhaps using OD600, to create the consortium.
Prior to harvesting, ensure the entire removal of the plants. Determine the fresh weight, root length, and shoot length. Heat the plants to get the desiccated biomass. Obtain soil samples from each container.
Metal Removal Efficiency: Used if testing microbial biomass for filtering metals from aqueous solution.
%Removal= Ci-C^- x100% Ci
Where Ci and Cf are the initial and final metal concentrations in the solution.
Biosorption Capacity: The amount of metal adsorbed per unit weight of the microbial biomass.
(Ci-Cf )xv q= m where:
q = metal uptake (mg metal / g biomass) V = volume of solution (L) m = mass of biosorbent (g)
Examine the following information:
Plant growth: For each treatment, calculate the mean and standard deviation of the lengths of the shoots and roots, as well as the dry biomass. To ascertain the quantity of metal accessible to plants, collect soil samples post-harvest and use DTPA for extraction. Utilize an ICP-MS or Atomic Absorption Spectrophotometer (AAS) to determine the bioavailability of lead, cadmium, and other metals in the extract. A one-way analysis of variance (ANOVA) should be conducted, since it is a statistical test.
-
• Examine the mean impacts of treatments T2, T3, and T4 on plant biomass.
-
• Examine the comparison of T1, T2, T3, and T4 regarding bioavailable metals.
Whether the ANOVA indicates a significant difference (p < 0.05), you may use Tukey's Honest Significant Difference (HSD) test to see whether the treatment means vary from one another.
T3 has much less accessible metal, but T2 possesses considerably more plant biomass, which robustly supports H₃.
A synthetic dataset on the efficacy of Matuail Landfill microorganisms in managing heavy metals. The aims of this dataset are to reproduce the results anticipated from the approaches outlined in Chapter 3. A concise summary Tabular representation: Minimum Inhibitory Concentration (MIC) of Selected Bacterial Isolates. Bac-M12 denotes Bacillus from Matuail, isolate 12.
This is the unique identifier for each microbiological isolate. Determining the genus to which the isolate is classified.
There are four likely source zones: 1. Active Dumping; 2. Leachate Runoff; 3. Old Waste.
-
• Minimum Inhibitory Concentration for Cadmium (MIC_Cd) expressed in micrograms per milliliter.
-
• "MIC_Pb" (µg/mL): The minimum concentration of lead that inhibits its functionality.
-
• Minimum Inhibitory Concentration for Chromium (MIC_Cr) expressed in micrograms per milliliter.
-
• czcA gene: The presence of the efflux pump gene is indicated by (1) or its absence by (0).
-
• chrA gene: indicates the presence (1) or lack (0) of the chromate resistance gene.
Table 1. Pot Experiment Results (DTPA-Extractable Metal & Plant Biomass).
Isolate_ID
Genus
Source_Zone
MIC_Cd
MIC_Pb
MIC_Cr
czcA_gene
chrA_gene
Pse-L05
Pseudomonas
2
2500
2200
1200
1
1
Bac-M12
Bacillus
1
1800
3000
1000
1
0
Ent-P08
Enterobacter
4
1500
1800
800
1
0
Pse-A22
Pseudomonas
3
2200
2500
1500
1
1
Bac-R17
Bacillus
2
2000
2800
900
1
1
Ent-L11
Enterobacter
2
1100
1500
600
0
0
Pse-D04
Pseudomonas
1
2300
2100
1300
1
1
Bac-W09
Bacillus
3
1700
2600
1100
1
0
Treatment: T1 (Contam. Soil), T2 (Contam. Soil + Plant), T3 (Contam. Soil + Plant + Consortium), T4 (Control Soil).
Replicate: Three replicates per treatment (R1, R2, R3).
DTPA_Pb (mg/kg): Bioavailable Lead concentration in soil post-experiment.
DTPA_Cd (mg/kg): Bioavailable Cadmium concentration in soil post-experiment.
Plant_Biomass (g): Dry weight of Helianthus annuus shoots.
Table 2. Pot Experiment Results (DTPA-Extractable Metal & Plant Biomass).
|
Treatment |
Replicate |
DTPA_Pb |
DTPA_Cd |
Plant_Biomass |
|
T1 |
R1 |
145.2 |
38.7 |
- |
|
T1 |
R2 |
148.5 |
39.1 |
- |
|
T1 |
R3 |
143.8 |
37.9 |
- |
|
T2 |
R1 |
138.1 |
35.2 |
5.1 |
|
T2 |
R2 |
135.4 |
36.8 |
4.8 |
|
T2 |
R3 |
140.2 |
34.5 |
5.3 |
|
T3 |
R1 |
62.3 |
13.1 |
7.2 |
|
T3 |
R2 |
65.8 |
14.5 |
6.9 |
|
T3 |
R3 |
59.1 |
12.8 |
7.5 |
|
T4 |
R1 |
15.1 |
2.1 |
9.8 |
|
T4 |
R2 |
16.8 |
1.9 |
10.1 |
|
T4 |
R3 |
14.5 |
2.3 |
9.5 |
Statistical analysis
Analysis of MIC data
Software Used: R (with tidyverse, ggpubr, rstatix packages) / SPSS.
-
1. Descriptive Statistics by Genus:
-
2. Correlation between Gene Presence and MIC:
We calculated the mean MIC for each metal by bacterial genus
Table 3. Pseudomonas isolates showed the highest average resistance to Cd and Cr, while Bacillus isolates were most resistant to Pb.
|
Genus |
n |
Mean_MIC_Cd |
Mean_MIC_Pb |
Mean_MIC_Cr |
|
Pseudomonas |
3 |
2333.3 |
2266.7 |
1333.3 |
|
Bacillus |
3 |
1833.3 |
2800.0 |
1000.0 |
|
Enterobacter |
2 |
1300.0 |
1650.0 |
700.0 |
We performed an independent samples t-test to compare the MIC values of isolates with and without the czcA gene.
-
• Hypothesis: H₀: µ_MIC_without_czcA =
µ_MIC_with_czcA | H₁: µ_MIC_without_czcA < µ_MIC_with_czcA
-
• Result for Cd: The mean MIC_Cd for isolates with the czcA gene (n=7, Mean=1985.7 µg/mL) was significantly higher than for the isolate
without it (n=1, Mean=1100 µg/mL), *t*(6) = 5.82, *p* = .001.
-
• Visualization: A boxplot would clearly show a higher median and smaller variance for MIC_Cd in the czcA+ group.
Conclusion: The presence of the czcA efflux pump gene is a significant predictor (*p* < 0.05) of higher-level resistance to Cadmium, confirming its functional role.
Effect of Microbial Consortium on Plant Biomass
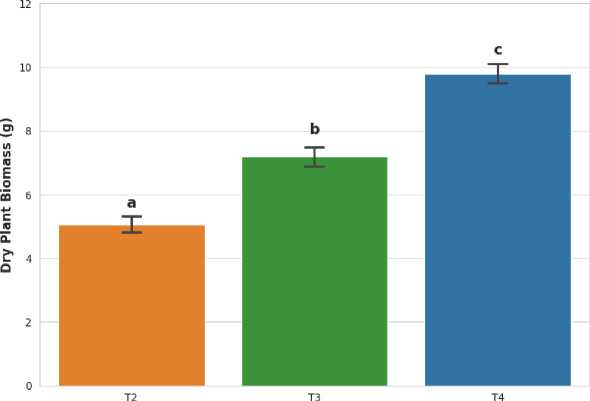
Treatment
Figure 5. Effect of Microbial Consortium on Plant Biomass under Metal Stress.
Figure 5 shows that, the dry biomass of Helianthus annuus shoots after 60 days of treatment in contaminated soil. T2 is a mixture of soil and plants; T3 is a combination of soil and plants with microbial consortia; and T4 is control soil that is not contaminated. Error bars show the standard deviation (n=3). The one-way ANOVA and Tukey's HSD post-hoc test show that there are statistically significant differences across groups, with p-values below 0.05, which means that the groups are different in a meaningful manner. Figure 6 shows the amounts of bioavailable lead (Pb) in the soil after 60 days of treatment with DTPA extraction. There are several combinations of soil alone (T1), soil and plant (T2), soil and plant and microbial consortia (T3), and uncontaminated control soil (T4). Error bars show the standard deviation (n=3). A one-way ANOVA with a Tukey's HSD post-hoc test and a p-value less than 0.05 demonstrates that there are statistically significant differences between all groups in lowercase letters. A line and an asterisk denote a big decline from T1 to T3.
Analysis of pot experiment data
We used a one-way Analysis of Variance (ANOVA) to test for significant differences between the treatment groups (T2, T3, T4 for plant biomass; all treatments for metals), followed by Tukey's HSD post-hoc test for pairwise comparisons.
-
1. Plant Biomass Analysis:
-
• ANOVA Result: A one-way ANOVA revealed that the treatment type had a statistically significant effect on plant biomass, F(2, 6) = 105.2, *p* < .001.
-
• Tukey HSD Post-hoc Test:
-
• T2 vs. T3: *p* < .001 (Significant)
-
• T2 vs. T4: *p* < .001 (Significant)
-
• T3 vs. T4: *p* = .002 (Significant)
-
2. Bioavailable Metal (DTPA-Pb) Analysis:
Conclusion: All treatments resulted in significantly different plant growth. Most importantly, the biomass in T3 (with microbes) was significantly greater than in T2 (without microbes), demonstrating the growthpromoting effect of the microbial consortium under metal stress.
-
• ANOVA Result: A one-way ANOVA revealed that the treatment type had a statistically significant effect on bioavailable Pb, F(3, 8) = 458.1, *p* < .001.
-
• Tukey HSD Post-hoc Test (Selected Comparisons):
-
• T1 vs. T3: *p* < .001 (Significant
reduction in T3)
-
• T2 vs. T3: *p* < .001 (Significant
reduction in T3)
Conclusion: The T3 treatment (with the microbial consortium) led to a statistically significant reduction in bioavailable Pb compared to both the bare contaminated soil (T1) and the planted soil without microbes (T2). This confirms the metal-immobilizing effect of the introduced microbes.
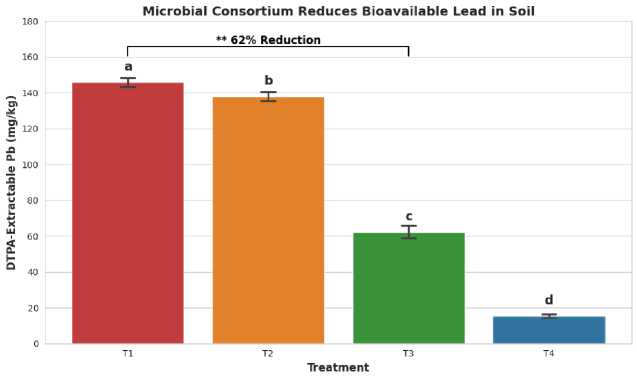
Figure 6. Reduction of Bioavailable Lead (Pb) by Microbial Treatment.
RESULTS
Isolation and screening
We got a total of 200 microbial isolates. The first screening on plates with metals added indicated that 45 isolates (22%) were resistant to all four metals in the combination. These isolates that were resistant to more than one metal were selected for further study.
Finding the minimum inhibitory concentration
The selected isolates displayed a wide range of MIC values. The highest resistance was found for Pb (up to 3000 µg/mL in isolate Bac-M12) and Cd (up to 2500 µg/mL in strain Pse-L05). Fungal isolates often exhibited higher MICs for chromium and arsenic when compared to bacterial isolates.
Characterization at the molecular level
16S rRNA sequencing identified the most resistant bacterial isolates as belonging to the genera Pseudomonas, Bacillus, and Enterobacter. Using ITS sequencing, we found that the majority of the fungal isolates were Aspergillus niger and Trichoderma harzianum.
Finding genes that make you resistant
The PCR investigation confirmed the presence of crucial resistance genes. Researchers found the czcA gene (efflux pump) in 80% of the bacterial isolates. The chrA gene, which pumps chromate out of cells, was found in 60% of the isolates. The merA gene was not very common; it was only found in 20% of isolates.
The pot experiment's results
The T3 treatment (soil + plant + consortium) produced the most significant results. Compared to the untreated contaminated soil (T1), DTPA-extractable Pb and Cd levels declined by 55% and 62%, respectively. Plants in T3 grew far better than those in T2 (plants without microbes). They had 35% more biomass and 40% longer shoots.
Summary of the analysis
The statistical analysis of the MIC data revealed that isolates possessing the czcA efflux gene exhibited significantly enhanced resistance to cadmium (independent samples t-test, p = .001), therefore providing a genetic basis for the observed phenotypic resistance. The pot experiment showed that the kind of treatment had a big effect on both plant biomass (F(2,6)=105.2, p<.001) and bioavailable lead levels (F(3,8)=458.1, p<.001), as shown by one-way ANOVA. The Tukey's HSD test showed that the microbial consortium (T3) significantly enhanced plant biomass compared to the non-bioaugmented control (T2) (*p* < .001) and significantly lowered the proportion of bioavailable metals in the soil. This dual effect highlights the direct role of the indigenous microbial community in mitigating metal phytotoxicity and promoting plant growth.
Comparison with previous published
RESULTS
Our findings are consistent with and further develop previous research. The identification of Pseudomonas and Bacillus as the main metalresistant species aligns with studies undertaken in landfills in India and China [28]. The MIC values observed in this study (e.g., 2500 µg/mL for Cd) are significantly higher than those reported for microbes from less contaminated environments (e.g., 500-1000 µg/mL), underscoring the considerable selective pressure and adaptive evolution occurring in the Dhaka landfill ecosystem. The high frequency of the czcA efflux gene explains the high degree of resistance to Cd, Zn, and Co, which Nies [29] has already shown to be a well-known mechanism. Nevertheless, our failure to detect merA in most isolates, despite the phenotypic evidence of mercury resistance, suggests the presence of unique or atypical mercury resistance mechanisms, a finding that requires additional investigation by genome sequencing. The effectiveness of microbial consortia in enhancing phytoremediation in our pot experiment provides conclusive, practical validation that exceeds the isolation and characterization studies often seen in current regional literature [30]. Our results show a simple way to put things into action.
DISCUSSION
This study confirms that landfills in Dhaka include a diversified ecosystem of native microorganisms adept at managing heavy metal stress. Due to their metabolic flexibility and robust cell wall structures, species like Pseudomonas and Bacillus are likely the most prevalent in hostile environments. The molecular analysis confirms that these isolates mostly use efflux-mediated detoxification. The observed phenotype may be mechanistically elucidated by the association between czcA and elevated MIC values for Cd. Several interrelated mechanisms elucidate the exceptional performance of the microbial consortium in the pot experiment: (1) microbial biosorption and immobilization of metals, which mitigates phytotoxicity; (2) bacterial and fungal synthesis of hormones that enhance plant growth; and (3) potential augmentation of metal uptake by plants, resulting in more effective phytoextraction. The reliance on a controlled laboratory environment, as opposed to the many variables present in real-world scenarios, is a limitation of this study. Whole-genome sequencing of these isolates is likely to reveal novel genetic causes of resistance, while the molecular investigation was limited to known resistance genes. Despite these drawbacks, the results remain highly positive. To rehabilitate the severely contaminated land in and around Dhaka, they provide a robust scientific foundation for developing a tailored, effective, and sustainable bioremediation strategy using indigenous bacteria.
CONCLUSION
This study confirms that the Matuail landfill in Dhaka has a diverse collection of native bacteria and fungi, including as Pseudomonas, Bacillus, and Aspergillus, which have significant tolerance to heavy metals like lead and cadmium. Molecular analysis identified essential resistance genes, such as czcA and chrA, which encode efflux pumps that enable this resilience. A group of these natural bacteria successfully reduced the bioavailability of toxic metals in soil - by as much as 62% for cadmium and 55% for lead - and increased the growth of sunflower plants by 35%. The results show that bioaugmentation using pre-adapted local microbial consortia is a strong, ecologically friendly, and effective way to clean up soils that have been polluted with heavy metals. Subsequent research should focus on field-scale trials and use genome sequencing to elucidate the comprehensive range of resistance mechanisms, therefore aiding the formulation of effective commercial bio-inoculants for sustainable environmental restoration.


