Обнаружение пневмонии на рентгеновских снимках грудной клетки с использованием сверточной нейронной сети на основе глубинного обучения
Автор: Мухриддин Араббоев, Шохрух Бегматов
Журнал: Современные инновации, системы и технологии.
Рубрика: Управление, вычислительная техника и информатика
Статья в выпуске: 5 (3), 2025 года.
Бесплатный доступ
Пневмония по-прежнему остается серьёзной проблемой общественного здравоохранения, особенно в условиях с ограниченными ресурсами, где доступ к квалифицированной радиологической диагностике затруднён. В данном исследовании предлагается метод на основе глубинного обучения с использованием собственной сверточной нейронной сети (CNN) для бинарной классификации рентгеновских снимков грудной клетки на категории «Пневмония» и «Нормал». Модель обучалась и оценивалась на специально подготовленном наборе данных, включающем 5 856 рентгеновских изображений грудной клетки, с применением методов предварительной обработки и увеличения данных для повышения обобщающей способности. Оценка предложенной CNN-модели показала высокие показатели эффективности: точность - 96,05%, точность (precision) — 98,79%, полнота (recall) - 95,76% и площадь под кривой ROC (AUC) - 0,9921. Кривая precision-recall продемонстрировала средний показатель точности 0,9970, что подтверждает устойчивость модели даже при наличии дисбаланса классов. Полученные результаты подчёркивают потенциал предложенной модели CNN как вспомогательного инструмента для быстрого и точного диагноза пневмонии, особенно в клинической практике и в условиях с ограниченными ресурсами.
Выявление пневмонии, рентгенография грудной клетки, глубинное обучение, сверточная нейронная сеть (CNN), классификация медицинских изображений, бинарная классификация, радиографическая диагностика, ROC-AUC, precision-recall, компьютерная диагностика (CAD).
Короткий адрес: https://sciup.org/14135194
IDR: 14135194 | DOI: 10.47813/2782-2818-2025-5-3-1018-1026
Текст статьи Обнаружение пневмонии на рентгеновских снимках грудной клетки с использованием сверточной нейронной сети на основе глубинного обучения
DOI:
Pneumonia is a potentially life-threatening respiratory infection caused by various pathogens, including bacteria, viruses, and fungi. It remains one of the leading causes of morbidity and mortality worldwide, particularly affecting young children, the elderly, and individuals with compromised immune systems. Early and accurate diagnosis of pneumonia is critical for effective treatment and improved patient outcomes.
Chest radiography (X-ray) is the most commonly employed imaging modality for pneumonia diagnosis due to its accessibility and speed. However, the interpretation of chest X-ray images requires specialized expertise and is susceptible to interobserver variability and human error, especially in resource-limited clinical settings.
Recent advancements in artificial intelligence (AI) and deep learning have shown great promise in automating the analysis of medical images. Convolutional Neural Networks (CNNs), in particular, have demonstrated exceptional performance in visual recognition tasks, including disease detection from radiographic images.
In this study, we propose a CNN-based approach for the binary classification of chest X-ray images into “Normal” and “Pneumonia” categories. A custom CNN architecture is designed and trained on a publicly available dataset, with performance evaluated using standard metrics such as accuracy, precision, recall, and AUC. The goal is to develop a reliable and computationally efficient model that can support clinicians in the rapid and accurate diagnosis of pneumonia.
The remainder of this paper is structured as follows. Section 2 reviews related work in the field of pneumonia detection using chest X-ray images, including both traditional and deep learning-based methods. It highlights key approaches, identifies limitations in existing models, and outlines the motivation for the proposed method. Section 3 provides a detailed overview of the dataset, including preprocessing steps and data augmentation strategies applied to improve model generalization. Section 4 introduces the architecture of the proposed Convolutional Neural Network (CNN), outlining its design principles and training configuration. Section 5 presents the experimental setup and performance evaluation, including metric definitions and comparative analysis. Section 6 discusses the results, highlighting strengths, limitations, and practical implications of the proposed model. Finally, Section 7 concludes the paper and outlines future directions for research and model deployment in clinical practice.
RELATED WORKS
In recent years, deep learning techniques -particularly Convolutional Neural Networks - have become central to medical image analysis [1]-[4], demonstrating strong performance in detecting pneumonia from chest X-ray images. Ayan and Ünver [5] employed the VGG16 architecture and achieved 84.5% classification accuracy, while Stephen et al. [6] proposed a conventional CNN model, reaching 93.73% accuracy. Jain et al. [7] leveraged transfer learning with VGG16 and VGG19, obtaining accuracies of 87.18% and 88.46%, respectively, highlighting the utility of pretrained models in extracting complex image features.
More recent studies have introduced enhanced architectures and ensemble methods. An et al. [8] combined EfficientNetB0 with DenseNet121 to achieve 95.19% accuracy and a precision of 98.38%. Similarly, Sharma and Guleria [9] integrated VGG16 with neural networks, reporting a balanced performance with 95.4% precision and recall. In 2025, studies by Walee et al. [10] and Shabaz et al. [11] showed that tailored CNNs with proper data augmentation strategies could reach accuracies exceeding 96%.
Research gap
Despite notable progress, many existing models suffer from limitations such as overfitting due to shallow architecture or poor generalization on unseen data. Several works lack extensive evaluation using robust metrics such as ROC-AUC or precision-recall curves, especially on well-augmented datasets. Furthermore, few studies have explored the performance of lightweight, custom CNNs that are optimized for deployment in real-world, low-resource healthcare settings.
Our contribution
To address these gaps, we propose a custom CNN architecture specifically designed for binary classification of chest X-ray images into “Pneumonia” and “Normal” categories. Unlike transfer learning approaches that rely on heavyweight pre-trained models, our model is built from scratch, offering a lightweight and efficient solution. It is trained on a curated and augmented dataset and rigorously evaluated using metrics including accuracy, precision, recall, ROC-AUC, and precision-recall curves. Our model achieves 96.05% accuracy, 98.79% precision, 95.76% recall, and an AUC of 0.9921, outperforming many existing approaches in terms of both accuracy and robustness.
MATERIALS AND METHODS
Dataset
The dataset used in this study comprises chest X-ray images classified into two categories: “Normal” (representing healthy individuals) and “Pneumonia” (representing patients diagnosed with pneumonia) [12]. The dataset is publicly available and widely used for benchmarking pneumonia detection models in the medical imaging domain.
Preprocessing and augmentation
To ensure consistency and improve model performance, all chest X-ray images were preprocessed prior to training. The original images were resized to a uniform resolution of 256 × 256 pixels, and pixel values were normalized to a range of [0, 1] to facilitate stable gradient descent during model optimization.
To enhance the generalization capability of the model and mitigate overfitting, data augmentation techniques were applied exclusively to the training set. Augmentation was performed using the ImageDataGenerator utility in Keras, incorporating the following random transformations:
-
• Rotation: Random rotation within a range of ±15 degrees
-
• Width and Height Shifts: Horizontal and vertical translations up to ±10% of the image dimensions
-
• Zoom: Random zooming within a ±10% range
-
• Horizontal Flipping: Random mirror flipping across the vertical axis
These augmentations simulate variations encountered in real-world clinical imaging scenarios, thus improving the model’s robustness and ability to learn discriminative features across diverse image conditions.
Data splitting
To ensure reliable model evaluation and maintain the integrity of class distribution, the dataset was divided into training and testing subsets using stratified sampling. This method preserves the proportion of “Normal” and “Pneumonia” cases in both subsets.
Specifically, 80% of the dataset was allocated for training, 10% for validation, and the remaining 10% for testing. This division allows the model to effectively learn discriminative features during training, while the validation and test sets enable an independent evaluation on unseen data, ensuring a reliable assessment of the model’s generalization performance.
MODEL ARCHITECTURE
A custom deep Convolutional Neural Network was developed from scratch to perform binary classification of chest X-ray images into “Normal” and “Pneumonia” categories. The proposed architecture is designed to balance model complexity with computational efficiency, enabling accurate feature extraction while maintaining practical training times.
The model consists of the following key components:
-
• Convolutional Blocks: Four sequential
convolutional blocks, each comprising a convolutional layer followed by a max pooling layer. The number of filters increases progressively across blocks: 32, 64, 128, and 256, enabling the network to capture both low- and high-level image features.
-
• Activation Function: Each convolutional layer uses the ReLU (Rectified Linear Unit) activation function to introduce non-linearity.
-
• Pooling Layers: MaxPooling2D layers are applied after each convolutional block to downsample the spatial dimensions, reducing computational load and mitigating overfitting.
-
• Flattening Layer: The final feature maps are flattened into a one-dimensional vector to interface with the dense layers.
-
• Fully Connected Layers: Two dense layers, each with 256 neurons, are used to learn high-level representations.
-
• Output Layer: A single neuron with a sigmoid activation function is used to produce the final binary classification output.
The model was compiled using the Adam optimizer and trained with the binary cross-entropy loss function. Performance was evaluated using standard metrics, including accuracy, precision, and recall, to assess classification effectiveness.
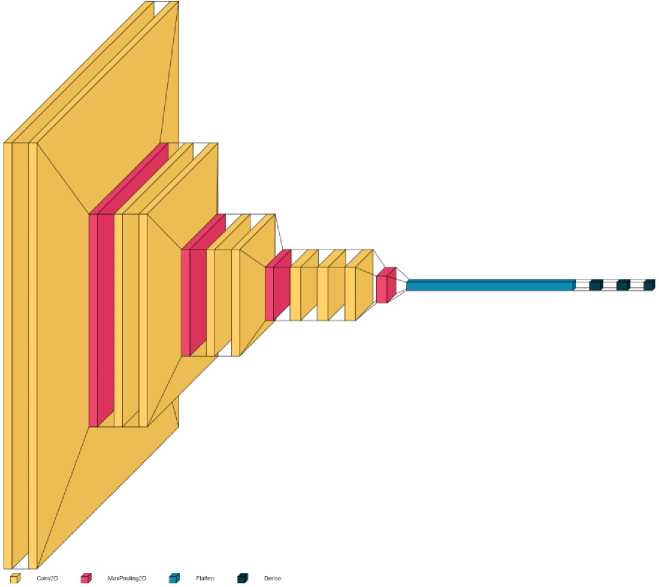
Figure 1. Architecture of the proposed deep convolutional neural network.
Figure 1 presents the architecture of the proposed Convolutional Neural Network model for pneumonia detection. The model comprises sequential Conv2D and MaxPooling2D layers for hierarchical feature extraction, followed by Flatten and Dense layers for final classification. This lightweight design ensures a balance between accuracy and computational efficiency.
Training strategy
The custom Convolutional Neural Network model was compiled with the Adam optimizer and binary cross-entropy loss. Evaluation metrics included accuracy, precision, and recall. The model was trained using mini-batches of 32 samples per iteration over a maximum of 50 epochs. To avoid overfitting and enable recovery of the best-performing model, two callback functions were integrated:
-
• EarlyStopping: Monitored validation loss with a patience of 5 epochs, restoring the best weights.
-
• ModelCheckpoint: Saved the best model during training based on validation performance.
The final model was saved and reloaded from the checkpoint containing the best validation performance for final evaluation.
Table 1 shows the hardware and software configuration used in the experiments, along with the dataset split proportions. These settings were adopted to ensure consistency and reproducibility of the training and evaluation process.
Table 1. System configuration and dataset split details
|
No. |
Name |
Value |
|
1 |
CPU of Computer system |
Intel Core i7-8700 |
|
2 |
RAM |
64 GB |
|
3 |
HDD |
4 TB |
|
4 |
Implementation tool |
Python, Pycharm |
|
5 |
Operating system |
Windows 10, 64 bit |
|
6 |
Training set |
80% data |
|
7 |
Validation set |
10% data |
|
8 |
Testing set |
10% data |
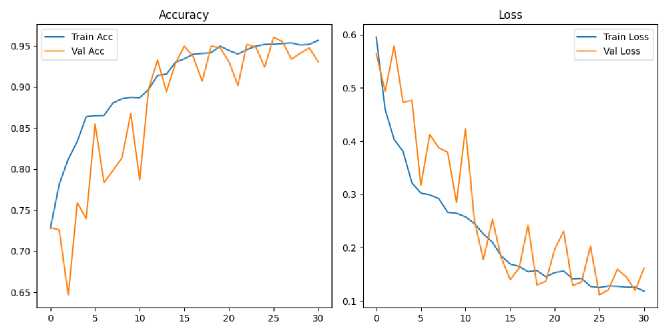
Figure 2. Training and validation accuracy and loss curves.
Figure 2 shows the training and validation accuracy (left) and loss (right) curves over 30 epochs for the proposed CNN model. The training accuracy shows a consistent upward trend, reaching over 96%, while validation accuracy remains stable above 93%, indicating good generalization. Similarly, both training and validation loss curves demonstrate a decreasing pattern, with no significant signs of overfitting. These trends confirm the model’s convergence and effective learning behavior during training.
EXPERIMENTAL RESULTS
Performance metrics
To evaluate the effectiveness of the proposed Convolutional Neural Network model, a comprehensive set of performance metrics was employed, including accuracy, precision, recall, and Area Under the Receiver Operating Characteristic Curve (AUC-ROC). These metrics provide a well-rounded assessment of the model’s ability to distinguish between pneumonia and normal chest X-ray images.
|
л (TP + TN) Accuracy =------------------ ) (TP + TN + FP + FN) |
(1) |
|
TP Precision =--------- (TP + FP) |
(2) |
|
TP Recall =---- TP--- (TP + FN) |
(3) |
In Equations (1-3), TP denotes True Positives, TN denotes True Negatives, FP denotes False Positives, and FN denotes False Negatives. These metrics are fundamental in evaluating the classification performance of the proposed CNN model.
To contextualize the model’s performance, Table 2 presents a comparative analysis with recent state-of-the-art studies on the same dataset. The proposed method outperformed many existing models in terms of classification accuracy and robustness.
Table 2. Comparative performance of the proposed CNN model with existing approaches for pneumonia detection
|
Ref. |
Year |
Dataset |
Classes |
Model |
Accuracy |
Precision |
Recall |
|
[5] |
2019 |
5856 |
2 |
VGG16 |
84.50 |
- |
- |
|
[6] |
2019 |
5856 |
2 |
CNN |
93.73 |
- |
- |
|
[7] |
2020 |
5840 |
3 |
VGG19 |
88.46% |
- |
95 |
|
[13] |
2022 |
1000 |
2 |
VGG16 |
95.07 |
- |
- |
|
[14] |
2022 |
5856 |
2 |
CNN |
96 |
- |
- |
|
[15] |
2022 |
5856 |
2 |
Quaternion CNN |
93.75 |
- |
- |
|
[16] |
2023 |
5863 |
2 |
CNN |
91 |
- |
- |
|
[17] |
2023 |
5856 |
2 |
NN with VGG16 |
95.4 |
95.4 |
95.4 |
|
[18] |
2024 |
5863 |
2 |
CNN |
59.9 |
77.75 |
59.9 |
|
[8] |
2024 |
5856 |
2 |
EfficientNetB0+Dens eNet121 |
95.19 |
98.38 |
93.84 |
|
[19] |
2025 |
5863 |
2 |
CNN |
92 |
- |
- |
|
[20] |
2025 |
5856 |
2 |
CNN |
90.22 |
- |
- |
|
[11] |
2025 |
5856 |
2 |
CNN |
96 |
93 |
96 |
|
[10] |
2025 |
5856 |
2 |
SVM |
93.5 |
94.7 |
95.80 |
|
This work |
5856 |
2 |
CNN |
96.05 |
98.79 |
95.76 |
|
A comprehensive comparison between the proposed Convolutional Neural Network model and existing state-of-the-art methods for pneumonia detection is presented in Table 2. The analysis highlights the effectiveness of the proposed approach in terms of accuracy, precision, and recall.
The proposed model achieved an accuracy of 96.05%, a precision of 98.79%, and a recall of 95.76%, thereby outperforming most of the referenced models. For instance, earlier studies employing the VGG16 architecture reported accuracy values ranging from 84.50% [5] to 95.07% [13], yet did not report corresponding precision or recall metrics, limiting the interpretability of their diagnostic robustness. Similarly, other models such as VGG19 [7] and Quaternion CNN [15] yielded accuracies of 88.46% and 93.75%, respectively - both lower than that of the proposed method.
Furthermore, although ensemble models such as EfficientNetB0 + DenseNet121 [8] achieved relatively strong performance (accuracy: 95.19%;
precision: 98.38%; recall: 93.84%), the proposed single CNN model surpassed them in all key evaluation metrics while maintaining architectural simplicity and reduced computational complexity.
Recent studies conducted in 2025 also report competitive results. For example, [11] achieved an accuracy and recall of 96.00%, but with a lower precision of 93.00%. Similarly, an SVM-based method [10] yielded a recall of 95.80%, comparable to the proposed model, but was limited by its lower overall accuracy (93.50%) and precision (94.70%).
These results demonstrate that the proposed CNN model not only matches or exceeds the diagnostic accuracy of more complex or ensemble-based systems but also maintains a higher degree of sensitivity (recall) and positive predictive value (precision). This makes it a suitable candidate for clinical deployment, especially in real-time or resource-limited environments where lightweight models are preferred.
Misclassified Chest X-ray images
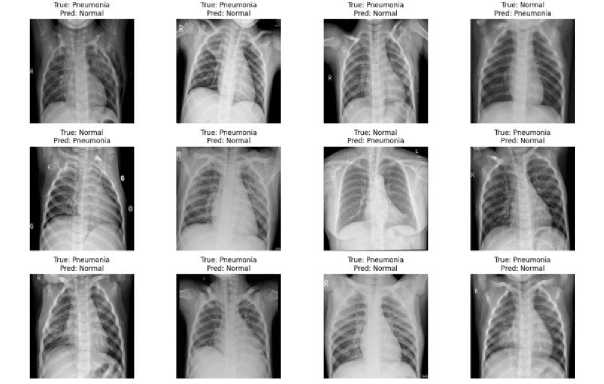
Figure 3. Misclassified chest X-ray images by the proposed model.
Figure 3 presents a selection of chest X-ray images that were incorrectly classified by the proposed model. The true and predicted labels are indicated for each image, showing instances where pneumonia cases were misclassified as normal and vice versa. These misclassifications highlight the challenges in differentiating between subtle radiographic features, underscoring the need for further model refinement or additional clinical input in ambiguous cases.
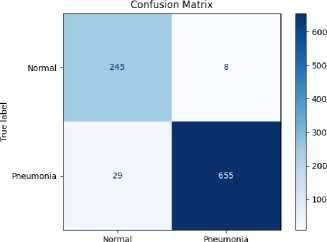
Predicted label
Figure 4. Confusion matrix of the proposed model for pneumonia detection.
Figure 4 depicts the confusion matrix representing the classification performance of the proposed model on the test set. The model correctly classified 245 normal and 655 pneumonia cases, while misclassifying 8 normal cases as pneumonia (false positives) and 29 pneumonia cases as normal (false negatives). The results demonstrate high accuracy, sensitivity, and specificity in distinguishing between normal and pneumonia-affected chest X-ray images.
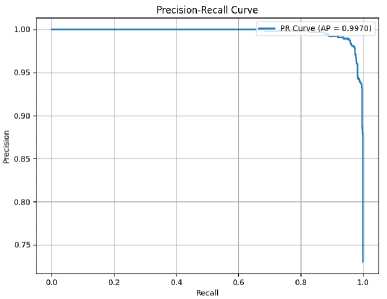
Figure 5. Precision-Recall curve of the proposed model with AP = 0.9970.
Figure 5 presents the Precision-Recall (PR) curve of the proposed model, which visualizes the trade-off between precision and recall across different classification thresholds. The high area under the curve (Average Precision, AP = 0.9970) indicates that the model maintains both high precision and high recall, even at varying thresholds. This reflects the model’s strong performance, particularly in scenarios with class imbalance where PR curves are more informative than ROC curves.
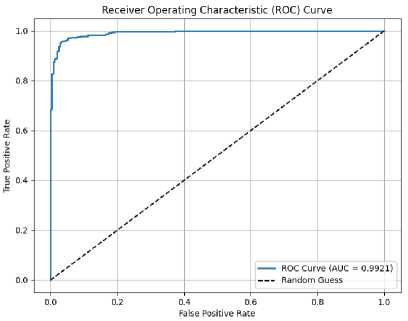
Figure 6. ROC curve of the proposed model with AUC = 0.9921.
Figure 6 illustrates the Receiver Operating Characteristic (ROC) curve of the proposed model, which demonstrates the trade-off between the true positive rate (sensitivity) and the false positive rate (1 – specificity). The curve shows that the model achieves a high level of classification performance, with an Area Under the Curve (AUC) of 0.9921. This indicates that the model has excellent discriminatory ability. For comparison, the diagonal dashed line represents the performance of a random classifier with an AUC of 0.5.
DISCUSSION
The experimental results demonstrate that the proposed custom Convolutional Neural Network (CNN) model offers strong and reliable performance for binary classification of chest X-ray images, effectively distinguishing between pneumonia and normal cases. With an achieved accuracy of 96.05%, precision of 98.79%, recall of 95.76%, and AUC of 0.9921, the model shows excellent potential as a diagnostic support tool in clinical settings.
The model’s high precision indicates its ability to minimize false positives, thereby reducing the likelihood of misclassifying healthy individuals as pneumonia patients. Similarly, the high recall suggests that the model is capable of correctly identifying the majority of pneumonia cases, which is crucial for timely intervention and treatment. The elevated AUC and average precision values further confirm the model’s robustness and reliability, even when dealing with potential class imbalance in the dataset.
The successful generalization to the test data can be attributed to multiple factors: the use of a well-curated dataset, application of diverse data augmentation techniques, and the implementation of training regularization strategies such as early stopping and model checkpointing. The architecture was designed to be both deep enough to capture complex radiographic features and computationally efficient for practical deployment.
However, certain limitations remain. The dataset used, while comprehensive, originates from a single source and may not fully represent the diversity of imaging conditions, equipment, or patient demographics encountered in real-world clinical environments. Additionally, the binary nature of the classification task restricts the model’s applicability to distinguishing pneumonia from other lung pathologies or subtypes such as bacterial versus viral pneumonia.
To further improve clinical relevance, future work should focus on external validation across multi-institutional datasets, incorporation of multi-class classification tasks, and integration with clinical decision support systems. Moreover, exploring lightweight model variants could enhance the applicability of the system in low-resource or mobile healthcare environments.
CONCLUSION
This study presents an effective deep learning approach for binary classification of chest X-ray images to detect pneumonia using a custom-designed Convolutional Neural Network. Through comprehensive preprocessing, augmentation, and model optimization strategies, the proposed model achieved impressive diagnostic performance, with an accuracy of 96.05%, precision of 98.79%, recall of 95.76%, and an AUC of 0.9921. The high average precision score (0.9970) further confirms the model’s ability to handle class imbalance and maintain consistent predictive performance.
These results demonstrate that the developed CNN is not only accurate but also computationally efficient, making it a strong candidate for deployment in real-time clinical decision support systems, particularly in resource-constrained settings. Future work will focus on expanding the model to multiclass pneumonia detection, validating it across external datasets, and integrating it into real-world medical workflows to assist radiologists in early and reliable diagnosis.


