Роль генов семейства Argonaute в эффектах активатора РНК-интерференции эноксацина на продолжительность жизни Drosophila melanogaster
Автор: Пакшина Н.Р., Яковлева Д.В., Уляшева Н.С., Прошкина Е.Н., Москалев А.А.
Журнал: Известия Коми научного центра УрО РАН @izvestia-komisc
Рубрика: Научные статьи
Статья в выпуске: 6 (64), 2023 года.
Бесплатный доступ
Эпигенетические механизмы играют ведущую роль в регуляции генной экспрессии и координации биологических процессов, влияя на скорость старения и продолжительность жизни организма. Важную роль в реализации этих механизмов играют малые РНК, которые подавляют активность своих мишеней путем РНК-интерференции и обеспечивают противовирусную защиту. Эноксацин является уникальным индуктором факторов РНК-интерференции с потенциальной геропротекторной активностью. Установлено, что его эффекты опосредованы микроРНК, но возможно участие и других видов некодирующих РНК. В данном исследовании мы изучили влияние эноксацина на продолжительность жизни Drosophila melanogaster и впервые проанализировали вклад в его эффект генов семейства Argonaute, которые специфично обеспечивают биогенез и функционирование микроРНК, киРНК и пивиРНК.
Малые рнк, рнк-интерференция, эноксацин, продолжительность жизни, старение, гены argonaute, drosophila melanogaster
Короткий адрес: https://sciup.org/149143620
IDR: 149143620 | УДК: 577.24 | DOI: 10.19110/1994-5655-2023-6-103-114
The role of genes of the Argonaute family in the effects of the RNA interference activator enoxacin on the lifespan of Drosophila melanogaster
Epigenetic mechanisms play a leading role in the regulation of gene expression and the coordination of biological processes, influencing the aging rate and the organism’s lifespan. An important role in the implementation of these mechanisms is played by small RNAs which suppress the activity of own targets through the RNA interference and provide the antiviral protection. Enoxacin is a unique inducer of RNA interference factors with potential geroprotective activity. Its effects have been identified to be mediated by miRNAs but other types of non-coding RNAs may also be involved. In this study, we have investigated the effect of enoxacin on the Drosophila melanogaster lifespan and first analyzed the contribution of Argonaute family genes to this effect which specifically ensure the biogenesis and functioning of miRNAs, siRNAs, and piRNAs.
Текст научной статьи Роль генов семейства Argonaute в эффектах активатора РНК-интерференции эноксацина на продолжительность жизни Drosophila melanogaster
Эпигенетика изучает наследуемые изменения экспрессии генов или клеточного фенотипа, не связанные с изменениями нуклеотидной последовательности генома. Эпигенетические механизмы включают в себя метилирование ДНК, модификацию РНК и гистонов, структуру хроматина и некодирующие РНК [1]. Предполагается, что нарушение их слаженной работы является одной из ведущих причин старения. Дисбаланс в эпигенетических механизмах вызывает обширные изменения генной экспрессии и состояние геномной нестабильности [2, 3].
Многочисленные исследования свидетельствуют о важной роли малых РНК в эпигенетической регуляции [1, 4, 5]. К данной группе некодирующих РНК относятся микроРНК, короткие интерферирующие РНК (киРНК), Piwi-взаимо- действующие РНК (пивиРНК), отличающиеся по размеру, функции и белкам Argonaute, с которыми они взаимодействуют. МикроРНК регулируют экспрессию генов с помощью РНК-интерференции - посттранскрипционного процесса путем нацеливания на специфические мРНК и последующего ингибирования трансляции и деградации этих молекул [1, 6]. Они участвуют в развитии организма, дифференцировке клеток, регуляции клеточного цикла, старении, метаболизме, апоптозе, и их экспрессия меняется при некоторых заболеваниях человека [1, 7]. КиРНК также осуществляют деградацию молекул-мишеней мРНК посредством РНК-интерференции. Кроме того, они участвуют в защите генома от активности мобильных генетических элементов и вирусов [8]. ПивиРНК описаны как важные регуляторы поддержания зародышевой линии. Они необходимы для подавления активности мобильных генетических элементов в зародышевых клетках [9], а также могут воздействовать на экспрессию генов посредством влияния на метилирование ДНК и модификации хроматина [10]. К белкам, обеспечивающим биогенез и функционирование малых РНК, относятся Drosha и представители семейства Dicer и Argonaute [11]. Имеются данные, указывающие на роль данных белков в регуляции стрессоустойчивости и продолжительности жизни (далее – ПЖ) модельных организмов [12].
Поиск терапевтических подходов, нацеленных на эпигенетические механизмы, является перспективной задачей современной биологии и медицины, так как изменения в этих механизмах имеют обратимый характер и тесно сопряжены с заболеваниями человека [13]. В настоящее время описан ряд веществ, обеспечивающих регуляцию метилирования ДНК, модификаций гистонов, транскрипционных регуляторов, которые способны влиять на скорость старения и предупреждать развитие возрастных патологических процессов [14-20]. Проводятся исследования, направленные на идентификацию низкомолекулярных соединений для ингибирования или активации экспрессии микроРНК [21, 22]. Эти молекулы также обладают потенциалом для замедления старения и предупреждения возраст-зависимых заболеваний [23–25].
Эноксацин является первым низкомолекулярным активатором факторов РНК-интерференции [21]. Данное соединение относится к семейству синтетических антибактериальных соединений на основе фторхинолонового скелета. Оно проявляет активность в отношении спектра грамотри-цательных и грамположительных бактерий [26]. Основной механизм его действия заключается в модификации процессинга микроРНК и усилении деградации мРНК посредством микроРНК и киРНК [21, 23, 27]. Но он также может влиять на функцию пивиРНК через микроРНК [28].
С использованием модели Drosophila melanogaster мы проверили геропротекторную активность эноксаци-на и оценили вклад конкретных путей биогенеза малых РНК в его эффект. Дрозофила в контексте данного исследования представляет собой уникальный модельный объект. У нее имеются белки семейства Argonaute, которые отвечают за биогенез конкретных типов малых РНК. Argonaute-1 (AGO1) необходим для синтеза и функционирования микроРНК, Argonaute-2 (AGO2) - для киРНК, бел- ки Argonaute-3 (AGO3), Aubergin (aub) и piwi осуществляют биогенез пивиРНК [29]. Для определения их роли мы сопоставили влияние эноксацина в разных концентрациях на ПЖ дрозофил линии дикого типа и дрозофил с нокдауном генов Argonaute, кодирующих эти белки.
Материалы и методы
Линии Drosophila melanogaster и получение особей с нокдауном генов Argonaute
Линии, использованные в работе, представлены в табл. 1. Они содержатся в коллекции лабораторных линий плодовых мушек Drosophila Института биологии ФИЦ Коми НЦ УрО РАН.
Условия содержания и обработки эноксацином
Для содержания дрозофил использовали климатические камеры KBF720-ICH (Binder, Германия). Животных содержали при температуре +25 °С, относительной влажности воздуха 60 %, 12-часовом режиме освещения. Состав питательной среды, на которой содержали контрольных и опытных животных при проведении экспериментов, был адаптирован из работы Xia и de Belle [30]: вода - 1 л, кукурузная мука – 92 г, сухие дрожжи – 32.1 г, агар-агар – 5.2 г, глюкоза - 136.9 г; для снижения микробиологической нагрузки 5 мл 10 %-ного раствора нипагина, разбавленного в 96 %-ном этаноле, 5 мл 50 %-ной пропионовой кислоты.
Раствор эноксацина в концентрациях 0.1, 0.5, 1, 5, 10, 50, 100, 500 мкг/мл наносили на поверхность питательной среды дрозофил в количестве 30 мкл на пробирку. В качестве растворителя использовали 1 мкмоль/л раствор NaOH. В контроле на среду наносили только 1 мкмоль/л NaOH.
Анализ продолжительности жизни
Для анализа ПЖ дрозофил собирали в течение 24 ч после вылета имаго. С использованием наркоза углекислым газом (Genesee Scientific, США) мух усыпляли, сортировали по полу и рассаживали в пробирки по 30 особей. Самцы и самки жили раздельно. Начиная с первого дня жизни имаго ежедневно вели подсчет числа умерших особей, два раза в неделю мух переносили на свежую среду.
Результаты представляли в виде кривых дожития и рассчитывали медианную ПЖ (длительность жизни наиболее типичных представителей выборки) и возраст 90 % смертности (показатель максимальной ПЖ). При статистической обработке данных применяли непараметрические методы, так как распределение продолжительности жизни не подчиняется нормальному закону. Для сравнения функций дожития использовали критерий Колмогорова-Смирнова [31]. Для оценки достоверности различий по медианной ПЖ – критерий Гехана-Бреслоу-Вилкоксона [32]. Для оценки статистической значимости различий максимальной ПЖ применяли метод Ванг-Аллисона [33]. Поправка Бонферрони использовалась для корректировки множественных сравнений [34]. Обработку данных проводили с помощью программы Statistica, версия 6.1 (StatSoft, США), статистической среды R, версия 2.15.1 (The R Foundation) и онлайн-приложения OASIS 2 (Online application for survival analysis) [35].
Линии Drosophila melanogaster , использованные для получения особей с РНК-интерференцией генов Argonaute
Drosophila melanogaster lines used to obtain individuals with RNA interference of the Argonaute genes
Table 1
|
Линии |
Генотип |
Описание |
Источник, номер линии |
|
Canton-S |
Линия дикого типа |
Центр линий дрозофил в Блумингтоне, США (#64349) |
|
|
P{CaryP} attP40 |
у[1] v[1]; P{y[+t7.7]=CaryP} Msp300[attP40] |
Контрольная линия для линий с РНК-интерференцией |
Центр линий дрозофил в Блумингтоне, США (#36304) |
|
P{CaryP} attP2 |
y[1] v[1]; P{y[+t7.7]=CaryP}attP2 |
Контрольная линия для линий с РНК-интерференцией |
Центр линий дрозофил в Блумингтоне, США (#36303) |
|
RNAi-AGO1 |
y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP. HM04006}attP2 |
Экспрессирует дцРНК для РНК-интерференции AGO1 под контролем промотора UAS в векторе VALIUM1. Конструкция в третьей хромосоме |
Центр линий дрозофил в Блумингтоне, США (#31700) |
|
RNAi-AGO2 |
y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS00108}attP2 |
Экспрессирует дцРНК для РНК-интерференции AGO2 под контролем промотора UAS в векторе VALIUM20. Конструкция в третьей хромосоме |
Центр линий дрозофил в Блумингтоне, США (#34799) |
|
RNAi-AGO3 |
y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP. HMC02938}attP40 |
Экспрессирует дцРНК для РНК-интерференции AGO3 под контролем промотора UAS в векторе VALIUM20. Конструкция во второй хромосоме |
Центр линий дрозофил в Блумингтоне, США (#44543) |
|
RNAi-piwi |
y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP. HMJ21827}attP40/CyO |
Экспрессирует дцРНК для РНК-интерференции piwi под контролем промотора UAS в векторе VALIUM20. Конструкция во второй хромосоме |
Центр линий дрозофил в Блумингтоне, США (#57819) |
|
RNAi-aub (1) |
y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP. JF01390}attP2 |
Экспрессирует дцРНК для РНК-интерференции aub под контролем промотора UAS в векторе VALIUM1. Конструкция во второй хромосоме |
Центр линий дрозофил в Блумингтоне, США (#31606) |
|
RNAi- aub (2) |
y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS00611}attP2 |
Экспрессирует дцРНК для РНК-интерференции aub под контролем промотора UAS в векторе VALIUM20. Конструкция в третьей хромосоме |
Центр линий дрозофил в Блумингтоне, США (#33728) |
|
GAL4-da |
w[*]; P{w[+mW.hs]=GAL4-da.G32}2; P{w[+mW.hs]=GAL4-da.G32}3a |
Повсеместная экспрессия GAL4 |
Центр линий дрозофил в Блумингтоне, США (#55849) |
Результаты и их обсуждение
Влияние эноксацина на продолжительность жизни особей Drosophila mela-nogaster дикого типа
Мы изучили влияние активатора РНК-интерференции эноксацина на ПЖ дрозофил линии дикого типа Canton-S . У самцов в двух биологических повторностях из трех (табл. 2) наблюдали увеличение медианной ПЖ дрозофил на 8-20 % (p < 0.05) и возраста 90 % смертности на 3–12 % (p < 0.05) при применении вещества в концентрациях 10-500 мкг/мл. У самок при этих же концентрациях также обнаружен положительный эффект. Медианная ПЖ повысилась на 5-8 % (p < 0.05), а возраст 90 % смертности - на 3-12 % (p < 0.05). На основании наших результатов можно говорить о геропротекторном потенциале эноксацина. Этот результат согласуется с данными других авторов, где показано, что эноксацин продлевает жизнь нематодам [21]. Тем не менее положительный эффект не всегда воспроизводился во всех биологических повторностях и при их совмещении был выражен слабо (табл. 2, рис. 1).
Влияние нокдауна генов Argonaute на эффект эноксацина
Для изучения вклада генов семейства Argonaute в эффект эноксацина мы изучили его влияние на ПЖ особей Drosophila melanogaster с нокдауном генов семейства Argonaute , кодирующих белки
Таблица 2
Параметры продолжительности жизни особей Drosophila melanogaster линии Canton-S при обработке эноксацином
Table 2
Lifespan parameters of Drosophila melanogaster Canton-S lines treated with enoxacin
|
Повторность |
C m |
Самцы |
Самки |
||||
|
M |
90 % |
N |
M |
90 % |
N |
||
|
1 |
Контроль |
63 |
68 |
136 |
68 |
76 |
143 |
|
1 |
63 |
69 |
143 |
68 |
76 |
152 |
|
|
5 |
63 |
68 |
142 |
69 |
76 |
142 |
|
|
10 |
60** |
68 |
143 |
68 |
76 |
147 |
|
|
50 |
61** |
67 |
148 |
64* |
76 |
156 |
|
|
100 |
61** |
68 |
141 |
64* |
74 |
152 |
|
|
500 |
63 |
68 |
144 |
64 |
76 |
146 |
|
|
2 |
Контроль |
50 |
60 |
144 |
60 |
67 |
136 |
|
1 |
49 |
60 |
144 |
60 |
68 |
112 |
|
|
5 |
53 |
63 |
125 |
61 |
69* |
133 |
|
|
10 |
53 |
60 |
144 |
63** |
71* |
143 |
|
|
50 |
54*** |
64* |
138 |
65*** |
75*** |
136 |
|
|
100 |
55* |
62* |
134 |
64* |
75** |
137 |
|
|
500 |
60*** |
67*** |
139 |
62 |
69 |
109 |
|
|
3 |
Контроль |
49 |
60 |
143 |
57 |
67 |
148 |
|
1 |
46 |
57 |
143 |
57 |
64 |
141 |
|
|
5 |
52 |
64 |
145 |
59 |
72 |
135 |
|
|
10 |
54*** |
66** |
145 |
61* |
71 |
148 |
|
|
50 |
51 |
64 |
137 |
60 |
68 |
141 |
|
|
100 |
50 |
61 |
134 |
58 |
67 |
137 |
|
|
500 |
42*** |
58 |
121 |
60* |
68 |
145 |
|
Примечания. Cm – концентрация эноксацина, мкг/мл; М – медианная ПЖ (сут); 90 % – возраст 90 % смертности (сут); N – количество особей в выборке.
Условные обозначения. * – различия с контролем статистически значимы при p < 0.05, ** – различия с контролем статистически значимы при p < 0.01, *** – различия с контролем статистически значимы при p < 0.001, достоверность различий для М указана по критерию Геха-на-Бреслоу-Вилкоксона с поправкой Бонферрони, достоверность различий для 90 % – по тесту Ванг-Аллисона с поправкой Бонферрони.
Note. Cm - concentration of enoxacin, µg/mL; M - median lifespan (days); 90 % - age of 90 % mortality (days); N - number of individuals in the sample.
Symbols. * - differences with control are statistically significant at p < 0.05; ** - differences with control are statistically significant at p < 0.01; *** - differences with control are statistically significant at p < 0.001, the significance of differences for M is indicated by the Gehan-Breslow Wilcoxon test with Bonferroni correction, the significance of differences for 90 % is indicated by the Wang-Allison test with Bonferroni correction.
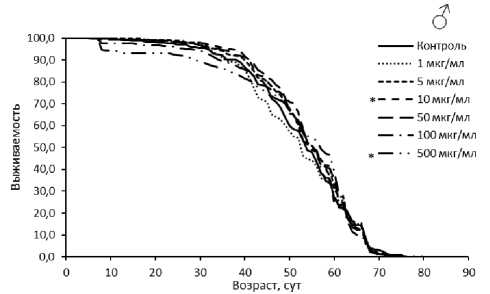
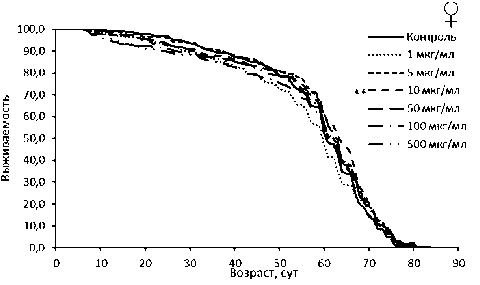
Рисунок 1. Кривые выживаемости самцов (А) и самок (Б) Drosophila melanogaster линии Canton-S при обработке эноксацином.
Условные обозначения. * p < 0.05, ** p < 0.01 – критерий Колмогорова-Смирнова с поправкой Бонферрони.
Figure 1. Survival curves for male (A) and female (Б) Canton-S Drosophila melanogaster treated with enoxacin.
Symbols. * p < 0.05, ** p < 0.01 – Kolmogorov-Smirnov test with Bonferroni correction.
биогенеза малых РНК, включая микроРНК ( AGO1 ), киРНК ( AGO2 ), пивиРНК ( AGO3, aub, piwi ).
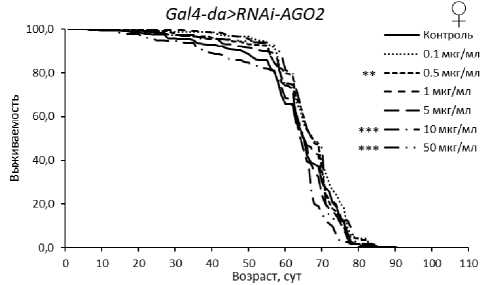
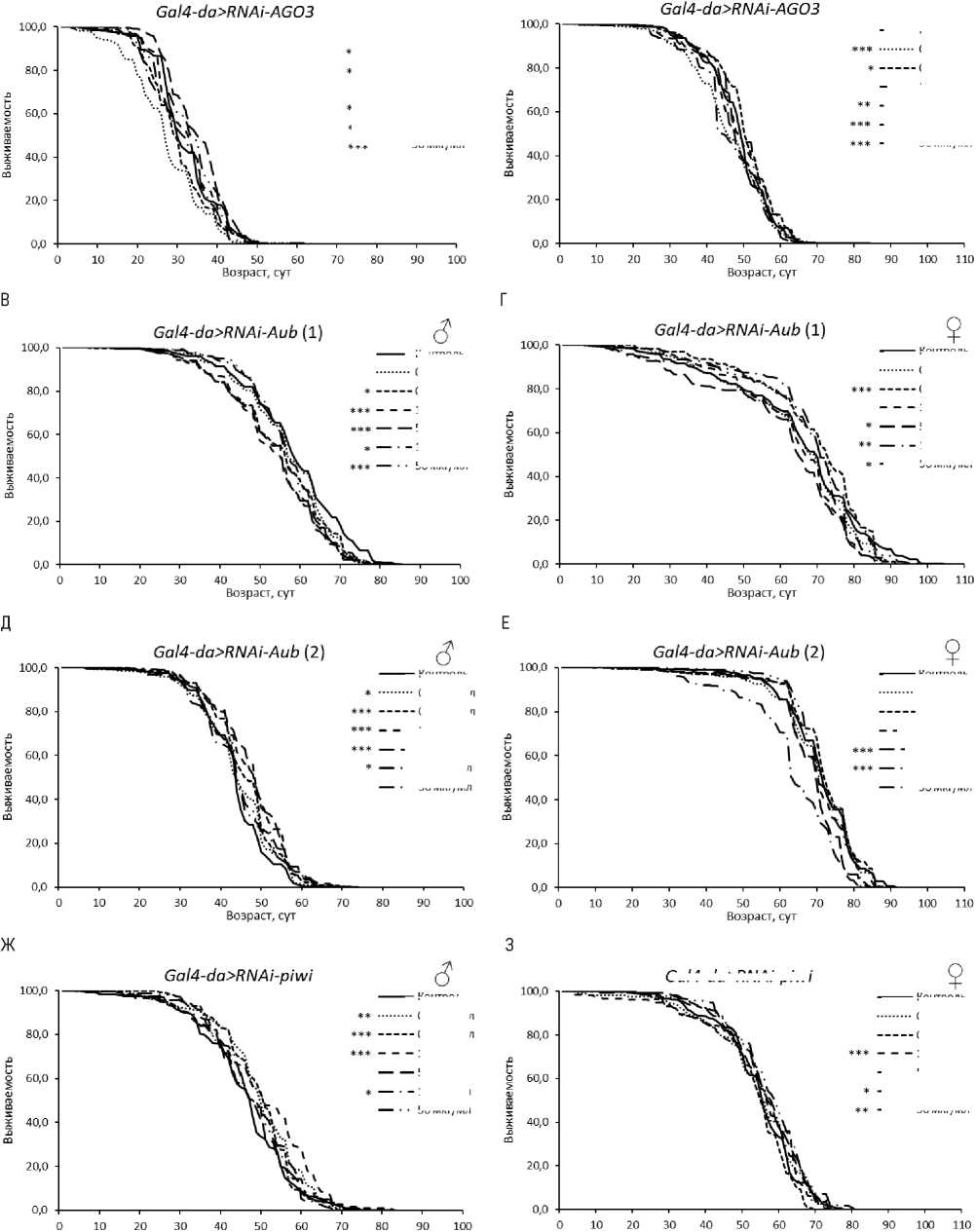
У самцов с нокдауном гена AGO2 обнаружен положительный эффект эноксацина в концентрациях 1, 10 и 50 мкг/мл в одной из повторностей (табл. 3, рис. 2). Также увеличение ПЖ при потреблении эноксацина наблюдали у мух с РНК-интерференцией aub® (при 0.5-50 мкг/мл вещества) и piwi (при 0.1-1 мкг/мл) (табл. 3, рис. 3). В перечисленных случаях медианная ПЖ была увеличена на 6–23 % (р < 0.05), а возраст 90 % смертности - на 5-23 % (р < 0.05). Эти данные указывают на то, что активность генов AGO2 , aub и piwi у самцов не снижает эффект эноксацина по сравнению с линией дикого типа.
У самцов с нокдауном AGO1, AGO3 и aub(1) эноксацин в изучаемых концентрациях либо имел менее выраженный и разнонаправленный эффект (между повторностями) на длительность жизни, либо воспроизводимо снижал медианную ПЖ на 3–13 % (p < 0.05) и возраст 90 % смертности на 5–17 % (p < 0.05). Данный результат говорит о возможном участии этих генов, отвечающих за биогенез и функционирование микроРНК и пивиРНК, в геропротекторном действии эноксацина.
У самок воспроизводимый положительный эффект энок-сацина наблюдался только при концентрации 0.5 мкг/мл у дрозофил с нокдауном aub(1) (р < 0.05). В остальных генотипах эноксацин либо укорачивал жизнь мух, либо не оказывал воспроизводимого влияния на изучаемые параметры ПЖ. Наибольшее снижение медианной ПЖ на 6–21 % (р < 0.05) и возраста 90 % смертности на 9-19 % (р < 0.05) при применении эноксацина обнаружено у дрозофил с РНК-интерференцией гена AGO1.
В то же время у дрозофил без нокдауна генов Argonaute наблюдали снижение параметров ПЖ на 1–35 % (p < 0.05) во всех исследуемых концентрациях.
Эноксацин является индуктором факторов РНК-интерференции, который нацелен, в первую очередь, на механизм биогенеза микроРНК [21, 23]. В исследованиях на клеточных культурах продемонстрировано, что это вещество способно усиливать опосредованную микроРНК деградацию мРНК и способствует биогенезу микроРНК и эндогенных киРНК [23]. В частности, эноксацин улучшает процессинг микроРНК путем связывания с TAR РНК-свя-зывающим белком 2 (TRBP) [21, 27]. Также он вовлекает Dicer совместно с AGO2 в процессинг предшественников микроРНК и способствует последующей загрузке регуляторных молекул в РНК-индуцированный комплекс сайленсинга (RISC) [23].
Эноксацин является многообещающим средством для лечения некоторых заболеваний, в том числе связанных со старением. Он ингибирует рост многих типов раковых клеток in vitro и in vivo, включая остеосаркому [36], меланому [37], рак предстательной железы [26], поджелудочной железы [38], легкого [39], щитовидной железы [40], шейки матки [41]. Кроме того, это вещество останавливает прогрессирование аутоиммунного процесса в тканях желчевыводящих путей [42], а также уменьшает вызванное диетой (с высоким содержанием жиров 60 %) ожирение у мышей, нормализует уровень глюкозы в крови и снижает симптомы бокового амиотрофического склероза [43, 44]. Эноксацин имеет низкий уровень токсичности, поскольку избирательно блокирует рост раковых клеток, не затрагивая здоровые клетки [45], безопасен для людей и широко применяется для лечения бактериальных инфекций мочевыводящих путей. Тем не менее на мышах было показано, что эноксацин не влияет на микробиоту кишечника (на содержание бактерий и распределение типов бактерий в кале) [43]. Дополнительно он оказывает противовирусное действие через усиление РНК-интерференции некоторых патогенных молекул с помощью киРНК, вплоть до потенциальной активности против SARS-CoV-2 [46-48].
В исследовании на нематодах Caenorhabditis elegans было показано, что эноксацин в концентрации 100 мкг/мл способен увеличивать ПЖ [21, 49]. На особях Drosophila melanogaster линии дикого типа Canton-S мы также показали, что эноксацин в концентрациях 10-500 мкг/мл способен увеличивать медианную и максимальную ПЖ до 20 %. Тем не менее этот эффект не всегда воспроизводился между биологическими повторностями. Более того, мы изучили эффекты эноксацина на трансгенных дрозофилах (но без нокдауна генов Argonaute ), содержащих конструкции P{CaryP}attP2 и P{CaryP}attP40 вместе с драйвером da-GAL4 . У этих мух наблюдали снижение ПЖ на 1–35 %
Параметры продолжительности жизни особей Drosophila melanogaster с нокдауном генов Argonaute при обработке эноксацином
Table 3
Lifespan parameters of Drosophila melanogaster specimens with Argonaute gene knockdown upon enoxacin treatment
|
Генотип |
Повторность |
C m |
Самцы |
Самки |
||||
|
M |
90% |
N |
M |
90% |
N |
|||
|
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
|
GAL4-da>RNAi-AGO1 |
1 |
Контроль |
49 |
60 |
145 |
64 |
71 |
156 |
|
0,1 |
46 |
57 |
142 |
70*** |
74* |
154 |
||
|
0,5 |
52** |
63 |
148 |
67*** |
73 |
142 |
||
|
1 |
52 |
61 |
149 |
65** |
71 |
154 |
||
|
5 |
53** |
63 |
150 |
64 |
71 |
152 |
||
|
GAL4-da>RNAi-AGO2 |
1 |
Контроль |
35 |
46 |
143 |
64 |
73 |
154 |
|
0,1 |
35 |
49 |
153 |
63 |
72 |
150 |
||
|
0,5 |
35 |
49 |
154 |
65 |
72 |
151 |
||
|
1 |
35 |
46 |
152 |
63 |
72 |
152 |
||
|
5 |
35 |
44 |
155 |
63 |
73 |
162 |
||
|
GAL4-da>RNAi-AGO3 |
1 |
Контроль |
28 |
35 |
148 |
49 |
57 |
139 |
|
0,1 |
24*** |
35* |
151 |
43*** |
57 |
151 |
||
|
0,5 |
26 |
37** |
154 |
49 |
56 |
139 |
||
|
1 |
27 |
40*** |
121 |
45 |
56 |
145 |
||
|
5 |
29** |
40 |
112 |
47 |
51* |
148 |
||
|
GAL4-da>RNAi-Aub (1) |
1 |
Контроль |
49 |
60 |
150 |
63 |
73 |
153 |
|
0,1 |
52*** |
64** |
158 |
59 |
73 |
155 |
||
|
0,5 |
49 |
63 |
156 |
64** |
78* |
152 |
||
|
1 |
46 |
57 |
153 |
63 |
77 |
144 |
||
|
5 |
49 |
64 |
154 |
63 |
73 |
147 |
||
|
GAL4-da>RNAi-Aub (2) |
1 |
Контроль |
42 |
53 |
148 |
71 |
78 |
144 |
|
0,1 |
43 |
56 |
137 |
66*** |
74** |
155 |
||
|
0,5 |
49*** |
56 |
146 |
70 |
78 |
158 |
||
|
1 |
45* |
56 |
151 |
65*** |
74*** |
156 |
||
|
5 |
49** |
57 |
157 |
70 |
77* |
156 |
||
|
GAL4-da>RNAi-piwi |
1 |
Контроль |
42 |
52 |
92 |
52 |
63 |
147 |
|
0,1 |
48* |
58* |
92 |
52 |
64 |
112 |
||
|
0,5 |
50*** |
57 |
93 |
54 |
63 |
118 |
||
|
1 |
47* |
64*** |
108 |
55* |
66 |
117 |
||
|
5 |
47*** |
61** |
138 |
54 |
64 |
51 |
||
|
Gal4-da > P{CaryP} attP40 |
1 |
Контроль |
70 |
78 |
161 |
70 |
78 |
159 |
|
0,1 |
66** |
77* |
162 |
70* |
78 |
168 |
||
|
0,5 |
70 |
78 |
163 |
63 |
77 |
167 |
||
|
1 |
67* |
77 |
160 |
64 |
77 |
162 |
||
|
5 |
65* |
77* |
162 |
67 |
77 |
155 |
||
|
Gal4-da>P{CaryP}attP2 |
1 |
Контроль |
58 |
72 |
150 |
46 |
74 |
152 |
|
0,1 |
53** |
65* |
150 |
37* |
77 |
154 |
||
|
0,5 |
55 |
72* |
161 |
57 |
77 |
156 |
||
|
1 |
58 |
72 |
153 |
53 |
78** |
153 |
||
|
5 |
53*** |
65*** |
155 |
74*** |
81*** |
150 |
||
|
GAL4-da>RNAi-AGO1 |
2 |
Контроль |
52 |
64 |
158 |
71 |
78 |
163 |
|
0,1 |
54 |
64 |
154 |
65** |
78 |
156 |
||
|
0,5 |
51 |
64 |
155 |
64*** |
71** |
158 |
||
|
1 |
47*** |
60** |
159 |
57*** |
64** |
155 |
||
|
5 |
46*** |
60 |
156 |
56*** |
63** |
158 |
||
|
10 |
48*** |
60 |
156 |
57*** |
68* |
159 |
||
|
50 |
53 |
63 |
158 |
68 |
75 |
62 |
||
|
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
|
GAL4-da>RNAi-AGO2 |
2 |
Контроль |
35 |
44 |
151 |
73 |
78 |
140 |
|
0,1 |
36 |
49 |
151 |
72 |
79 |
159 |
||
|
0,5 |
36 |
44 |
154 |
70 |
78 |
154 |
||
|
1 |
38* |
48 |
150 |
70 |
77 |
155 |
||
|
5 |
35 |
47 |
145 |
68*** |
77 |
164 |
||
|
10 |
38 |
50* |
151 |
67*** |
74 |
156 |
||
|
50 |
37** |
50** |
153 |
71 |
79 |
156 |
||
|
GAL4-da>RNAi-AGO3 |
2 |
Контроль |
37 |
46 |
86 |
52 |
64 |
149 |
|
0,1 |
33*** |
42*** |
84 |
51** |
60* |
149 |
||
|
0,5 |
34* |
44 |
83 |
54 |
64 |
153 |
||
|
1 |
36* |
42* |
127 |
52 |
61 |
158 |
||
|
5 |
39 |
46 |
141 |
54 |
62*** |
164 |
||
|
10 |
36*** |
40*** |
136 |
53** |
60*** |
157 |
||
|
50 |
36 |
44 |
147 |
49*** |
60*** |
152 |
||
|
GAL4-da>RNAi-Aub (1) |
2 |
Контроль |
64 |
74 |
152 |
71 |
81 |
157 |
|
0,1 |
64 |
72 |
152 |
78*** |
86 |
155 |
||
|
0,5 |
64 |
72 |
162 |
79*** |
86 |
155 |
||
|
1 |
61 |
71 |
187 |
72 |
81 |
153 |
||
|
5 |
60** |
70*** |
159 |
71 |
82 |
144 |
||
|
10 |
60** |
69 |
148 |
71 |
82 |
157 |
||
|
50 |
60*** |
68** |
162 |
68 |
78 |
147 |
||
|
GAL4-da>RNAi-Aub (2) |
2 |
Контроль |
45 |
57 |
154 |
79 |
86 |
144 |
|
0,1 |
45 |
58 |
149 |
79 |
86* |
156 |
||
|
0,5 |
46 |
58 |
165 |
78 |
86*** |
154 |
||
|
1 |
49** |
57 |
151 |
78 |
85*** |
157 |
||
|
5 |
50*** |
61* |
155 |
71*** |
78*** |
157 |
||
|
10 |
47 |
60*** |
145 |
71*** |
82*** |
128 |
||
|
50 |
47 |
60* |
154 |
74*** |
82*** |
156 |
||
|
GAL4-da>RNAi-piwi |
2 |
Контроль |
47 |
61 |
73 |
63 |
72 |
116 |
|
0,1 |
57* |
65 |
80 |
61* |
69 |
106 |
||
|
0,5 |
53 |
62 |
76 |
60*** |
67*** |
81 |
||
|
1 |
58*** |
68 |
72 |
60** |
70* |
95 |
||
|
5 |
51 |
61 |
68 |
59** |
70 |
124 |
||
|
10 |
46 |
60 |
89 |
64 |
74 |
110 |
||
|
50 |
50 |
60 |
100 |
63 |
69 |
148 |
||
|
Gal4-da > P{CaryP} attP40 |
2 |
Контроль |
64 |
75 |
65 |
79 |
150 |
|
|
0,1 |
57*** |
68*** |
156 |
57** |
75* |
163 |
||
|
0,5 |
57*** |
70*** |
158 |
60 |
79 |
156 |
||
|
1 |
60** |
70 |
159 |
49*** |
77 |
142 |
||
|
5 |
51*** |
68 |
154 |
49*** |
68*** |
152 |
||
|
10 |
56*** |
67*** |
117 |
42*** |
67*** |
154 |
||
|
50 |
57*** |
67*** |
152 |
60*** |
68*** |
156 |
||
|
Gal4-da>P{CaryP}attP2 |
2 |
Контроль |
50 |
65 |
155 |
80 |
89 |
153 |
|
0,1 |
51 |
64 |
157 |
78* |
86* |
138 |
||
|
0,5 |
51 |
64 |
152 |
74*** |
86*** |
155 |
||
|
1 |
44** |
64 |
168 |
70*** |
83*** |
158 |
||
|
5 |
47 |
64 |
186 |
56*** |
79*** |
152 |
||
|
10 |
54 |
64 |
148 |
64*** |
76*** |
138 |
||
|
50 |
48 |
64 |
161 |
67*** |
82*** |
160 |
||
|
GAL4-da>RNAi-AGO1 |
3 |
Контроль |
49 |
60 |
157 |
67 |
78 |
142 |
|
10 |
50 |
58 |
161 |
63** |
74 |
153 |
||
|
50 |
51** |
63 |
164 |
66 |
74 |
160 |
||
|
GAL4-da>RNAi-AGO2 |
3 |
Контроль |
31 |
44 |
158 |
59 |
73 |
149 |
|
10 |
32 |
44 |
156 |
63 |
71 |
155 |
||
|
50 |
32 |
44 |
159 |
65*** |
78* |
159 |
|
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
|
GAL4-da>RNAi-AGO3 |
3 |
Контроль |
30 |
43 |
157 |
48 |
52 |
157 |
|
10 |
27*** |
41* |
157 |
43*** |
55 |
162 |
||
|
50 |
27*** |
41* |
161 |
49*** |
57* |
160 |
||
|
GAL4-da>RNAi-Aub (1) |
3 |
Контроль |
65 |
78 |
160 |
78 |
94 |
160 |
|
10 |
57*** |
69*** |
157 |
71*** |
85*** |
159 |
||
|
50 |
57*** |
65*** |
161 |
78 |
87 |
154 |
||
|
GAL4-da>RNAi-Aub (2) |
3 |
Контроль |
44 |
51 |
159 |
67 |
78 |
160 |
|
10 |
44* |
56 |
159 |
63*** |
73 |
159 |
||
|
50 |
44 |
53 |
157 |
70*** |
80** |
156 |
||
|
GAL4-da>RNAi-piwi |
3 |
Контроль |
48 |
62 |
147 |
55 |
63 |
145 |
|
10 |
50 |
60 |
160 |
55 |
63 |
154 |
||
|
50 |
47 |
60 |
155 |
56 |
68* |
167 |
||
|
Gal4-da > P{CaryP} attP40 |
3 |
Контроль |
63 |
71 |
155 |
56 |
78 |
153 |
|
10 |
56*** |
64*** |
157 |
51** |
70** |
153 |
||
|
50 |
57*** |
64* |
145 |
64 |
78 |
160 |
||
|
Gal4-da>P{CaryP}attP2 |
3 |
Контроль |
53 |
65 |
161 |
78 |
91 |
147 |
|
10 |
45*** |
57*** |
157 |
78*** |
81*** |
155 |
||
|
50 |
49*** |
63** |
160 |
80 |
88 |
160 |
Примечания. Cm – концентрация эноксацина, мкг/мл; М – медианная ПЖ (сут); 90 % – возраст 90 % смертности (сут). N – количество особей в выборке. Условные обозначения. * – различия с контролем статистически значимы при p < 0.05; ** – различия с контролем статистически значимы при p < 0.01, *** – различия с контролем статистически значимы при p < 0.001, достоверность различий для М указана по критерию Гехана-Бреслоу-Вилкоксона с поправкой Бонферрони, достоверность различий для 90 % указана по тесту Ванг-Аллисона с поправкой Бонферрони.
Note. Cm - concentration of enoxacin µg/mL; M - median lifespan (days); 90 % - age of 90 % mortality (days); N - number of individuals in the sample. Symbols. * - differences with control are statistically significant at p < 0.05; ** - differences with control are statistically significant at p < 0.01; *** -differences with control are statistically significant at p < 0.001, the significance of differences for M is indicated by the Gehan-Breslow Wilcoxon test with Bonferroni correction, the significance of differences for 90 % is indicated by the Wang-Allison test with Bonferroni correction.
Б
А
В
Г
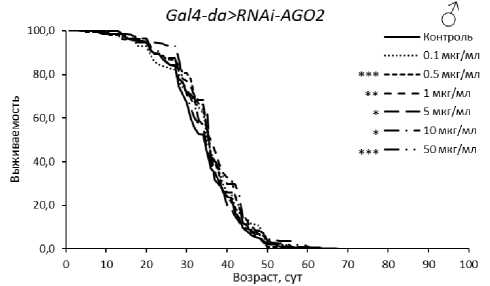
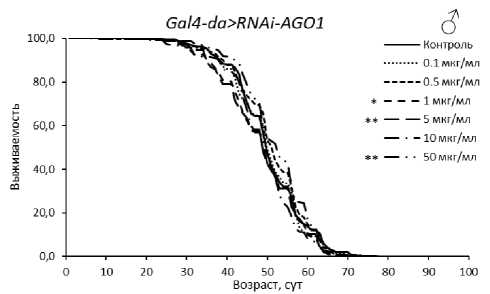
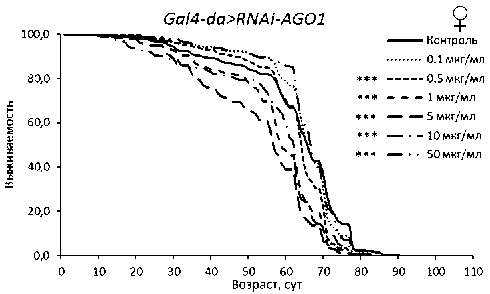
Рисунок 2. Кривые выживаемости особей Drosophila melanogaster с нокдауном генов подсемейства Argonaute при обработке эноксацином.
Условные обозначения. *p < 0.05; **p < 0.01; ***p < 0.001 – критерий Колмогорова-Смирнова с поправкой Бонферрони.
Figure 2. Survival curves of Drosophila melanogaster individuals with Argonaute subfamily gene knockdown treated with enoxacin.
Symbols. *p < 0.05; **p < 0.01; ***p < 0.001 – Kolmogorov-Smirnov test with Bonferroni correction.
Gal4-da>RNAi-piwi
■ - Контроль
0.1 мкг/мл 0.5 мкг/мл — — 1 мкг/мл — 5 мкг/мл • -10 мкг/мл - - 50 мкг/мл
" - Контроль
0.1 мкг/мл 0.5 мкг/мл 1 мкг/мл 5 мкг/мл 10 мкг/мл — ■ • 50 мкг/мл
Возраст, сут
Возраст, сут
Рисунок 3. Кривые выживаемости особей Drosophila melanogaster с нокдауном генов подсемейства PIWI при обработке эноксацином.
Условные обозначения. *p < 0.05; **p < 0.01; ***p < 0.001 – критерий Колмогорова-Смирнова с поправкой Бонферрони.
Figure 3. Survival curves of Drosophila melanogaster individuals with PIWI subfamily gene knockdown treated with enoxacin.
Symbols. *p < 0.05; **p < 0.01; ***p < 0.001 – Kolmogorov-Smirnov test with Bonferroni correction.
------ Контроль *++ ......... 0.1 мкг/мл
*** -----о.5мкг/мл ** — — — 1 мкг/мл *** — ^5 мкг/мл *** — . —10 мкг/мл *** — • • 50 мкг/мл
11 - Контроль
0.1 мкг/мл
0.5 мкг/мл
— — 1 мкг/мл — — 5 мкг/мл — ■ - 10 мкг/мл — ■ • 50 мкг/мл
Контроль 0.1 мкг/мл 0.5 мкг/мл 1 мкг/мл 5 мкг/мл 10 мкг/мл 50 мкг/мл
Контроль 0.1 мкг/мл 0.5 мкг/мл — 1 мкг/мл — 5 мкг/мл ■ -10 мкг/мл ■ • 50 мкг/мл
Контроль 0.1 мкг/мл 0.5 мкг/мл 1 мкг/мл 5 мкг/мл 10 мкг/мл 50 мкг/мл
■ Контроль
0.1 мкг/мл
0.5 мкг/мл 1 мкг/мл
— — 5 мкг/мл — - - 10 мкг/мл — - • 50 мкг/мл
при всех концентрациях индуктора РНК-интерференции. Это говорит о том, что действие эноксацина на старение и ПЖ организма может зависеть от комплекса внешних и внутренних факторов, и требуется детальное изучение лежащих в его основе механизмов.
Согласно результатам работы на нематоде, эноксацин действует на продолжительность жизни и старение через пути митогормезиса и SKN-1/Nrf2, снижая при этом уровень miR-34-5p [21,49]. Также в качестве мишени данного соединения у нематод описан РНК-специфическая аденозиндеаминаза (ADAR). При утрате функции ADAR у червей исчезал положительный эффект применения эноксацина [49]. Тем не менее, ввиду специфики организации аппарата биогенеза малых РНК у червей, до конца неясно, связан ли механизм действия эноксацина только с микроРНК, либо он также опосредован функционированием киРНК и пивиРНК.
У Drosophila melanogaster есть пять генов семейства Argonaute , относящиеся к двум подсемействам - Argonaute (AGO1, AGO2) и PIWI (AGO3, aub, piwi) , которые играют важную роль в регуляции экспрессии генов и транспозонов. Мы оценили вклад конкретных генов Argonaute , специфичных для разных типов малых РНК молекул, в эффект эноксацина на длительность жизни дрозофил. Ген AGO1 повсеместно экспрессируется в ходе развития, его белок обеспечивает активность связывания микроРНК, регулирует экспрессию генов, подавляя трансляцию [50]. AGO2 также повсеместно экспрессируется, а его белок выполняет функцию защиты от транспозонов и вирусов путем связывания с киРНК и участвует в формировании комплекса RISC [51]. Белки подсемейства PIWI (AGO3, Aub, piwi) необходимы для репрессии транспозонов зародышевой линии, однако ген piwi экспрессируется также в соматических клетках гонад дрозофилы [50–52].
В данном исследовании мы обнаружили, что у особей Drosophila melanogaster с нокдауном гена AGO1 эноксацин имел либо отрицательный, либо слабый положительный эффект на ПЖ. Полученный результат указывает на вклад механизма биогенеза микроРНК в спектр биологических активностей этого соединения, что согласуется с указанными выше литературными данными, полученными на клеточных культурах и нематодах.
Дрозофилы с РНК-интерференцией гена AGO2 реагировали на эноксацин схожим образом с линией дикого типа Canton-S . Эноксацин оказывал либо положительное действие, либо не вызывая статистически значимых изменений ПЖ. По-видимому, механизм биогенеза киРНК в меньшей степени определяет геропротекторную активность данного вещества, что согласуется с результатами анализа, проведенного на нематодах [49].
Особи с нокдауном генов подсемейства PIWI демонстрировали неожиданные эффекты эноксацина на ПЖ. В ряде случаев эноксацин вызывал снижение ПЖ у дрозофил с нокдауном AGO3, aub и piwi, что может указывать на вклад пивиРНК и генов PIWI в эффекты эноксацина и детерминирование жизнеспособности взрослого организма в целом. Этот вопрос требует детального изучения. В первую очередь, в связи с тем, что в настоящее время предполагается, что решающую роль пивиРНК имеет в за- родышевой линии и половых клетках [53, 54]. Тем не менее, в работе на раковых клетках человека было показано, что эноксацин может восстанавливать активность PIWIL3 (представитель подсемейства белков PIWI, обеспечивающих выработку пивиРНК) через микроРНК [28].
Список литературы Роль генов семейства Argonaute в эффектах активатора РНК-интерференции эноксацина на продолжительность жизни Drosophila melanogaster
- Yao, Q. The roles of microRNAs in epigenetic regulation / Q. Yao, Y. Chen, X. Zhou // Curr Opin Chem Biol. – 2019. – Vol. 51. – P. 11-17.
- Yang, J. H. Loss of epigenetic information as a cause of mammalian aging / J. H. Yang, M. Hayano, P. T. Griffin [et al.] // Cell. – 2023. – Vol. 186, № 2. – P. 305-326.e27.
- Sen, P. Epigenetic mechanisms of longevity and aging / P. Sen, P. P. Shah, R. Nativio [et al.] // Cell. – 2016. – Vol. 166, № 4. – P. 822-839.
- Huang, X. A. A major epigenetic programming mechanism guided by piRNAs / X. A. Huang, H. Yin, S. Sweeney [et al.] // Dev Cell. – 2013. – Vol. 24, № 5. – P. 502-516.
- Duempelmann, L. Small RNAs in the transgenerational inheritance of epigenetic information / L. Duempelmann, M. Skribbe, M. Bühler // Trends Genet. – 2020. – Vol. 36, № 3. – P. 203-214.
- Sankrityayan, H. Diabetic nephropathy: The regulatory interplay between epigenetics and microRNAs / H. Sankrityayan, Y. A. Kulkarni, A. B. Gaikwad // Pharmacol Res. – 2019. – Vol. 141. – P. 574-585.
- Iorio, M.V. Interplay between microRNAs and the epigenetic machinery: an intricate network / M. V. Iorio, C. Piovan, C. M. Croce // Biochim Biophys Acta. – 2010. – Vol. 1799, № 10-12. – P. 694-701.
- Moazed, D. Small RNAs in transcriptional gene silencing and genome defence / D. Moazed // Nature. – 2009. – Vol. 457, № 7228. – P. 413-420.
- Peters, L. Argonaute proteins: mediators of RNA silencing / L. Peters, G. Meister // Mol Cell. – 2007. – Vol. 26, № 5. – P. 611-623.
- Jia, D. D. The regulatory function of piRNA/PIWI complex in cancer and other human diseases: The role of DNA methylation / D. D. Jia, H. Jiang, Y. F. Zhang [et al.] // Int J Biol Sci. – 2022. – Vol. 18, № 8. – P. 3358-3373.
- Kim, V.N. Biogenesis of small RNAs in animals / V.N. Kim, J. Han, M.C. Siomi // Nat. Rev. Mol. Cell Biol. – 2009. – Vol. 10. – P. 126-139.
- Proshkina, E. N. Genome-Protecting Compounds as Potential Geroprotectors / E. N. Proshkina, M. V. Shaposhnikov, A. A. Moskalev // Int J Mol Sci. – 2020. – Vol. 21, № 12. – P. 4484.
- Memari, F. Epigenetics and Epi-miRNAs: Potential markers/ therapeutics in leukemia / F. Memari, Z. Joneidi, B. Taheri [et al.] // Biomed Pharmacother. – 2018. – Vol. 106. – P. 1668-1677.
- Cheng, Y. Targeting epigenetic regulators for cancer therapy: mechanisms and advances in clinical trials / Y. Cheng, C. He, M. Wang [et al.] // Signal Transduct Target Ther. – 2019. –Vol. 4. – P. 62.
- Audia, J. E. Histone modifications and cancer / J. E. Audia, R. M. Campbell // Cold Spring Harb Perspect Biol. – 2016. – Vol. 8, № 4. – P. a019521.
- Yang, X. Targeting DNA methylation for epigenetic therapy / X. Yang, F. Lay, H. Han [et al.] // Trends Pharmacol Sci. – 2010. – Vol. 31, № 11. – P. 536-546.
- Siklos, M. Therapeutic targeting of chromatin: status and opportunities / M. Siklos, S. Kubicek // FEBS J. – 2022. – Vol. 289, № 5. – P. 1276-1301.
- Morera, L. Targeting histone methyltransferases and demethylases in clinical trials for cancer therapy / L. Morera, M. Lübbert, M. Jung // Clin Epigenetics. – 2016. – Vol. 8. – P. 57.
- Dai, E. Epigenetic modulation of antitumor immunity for improved cancer immunotherapy / E. Dai, Z. Zhu, S. Wahed [et al.] // Mol Cancer. – 2021. – Vol. 20, № 1. – P. 171.
- Cao, J. Cancer epigenetics, tumor immunity, and immunotherapy / J. Cao, Q. Yan // Trends Cancer. – 2020. – Vol. 6, № 7. – P. 580-592.
- Felicetti, T. Modulating microRNA processing: enoxacin, the progenitor of a new class of drugs / T. Felicetti, V. Cecchetti, G. Manfroni // J Med Chem. – 2020. – Vol. 63, № 21. – P. 12275-12289.
- Zhang, S. Targeting microRNAs with small molecules: from dream to reality / S. Zhang, L. Chen, E. J. Jung [et al.] // Clin Pharmacol Ther. – 2010. – Vol. 87, № 6. – P. 754-758.
- Shan, G. A small molecule enhances RNA interference and promotes microRNA processing / G. Shan, Y. Li, J. Zhang [et al.] // Nat Biotechnol. – 2008. – Vol. 26, № 8. – P. 933-940.
- Zhao, R. Designing strategies of small-molecule compounds for modulating non-coding RNAs in cancer therapy / R. Zhao, J. Fu, L. Zhu [et al.] // J Hematol Oncol. – 2022. – Vol. 15, № 1. – P. 14.
- Wang, K. Epigenetic regulation of aging: implications for interventions of aging and diseases / K. Wang, H. Liu, Q. Hu [et al.] // Signal Transduct Target Ther. – 2022. – Vol. 7, № 1. – P. 374.
- Sousa, E. Enoxacin inhibits growth of prostate cancer cells and effectively restores microRNA processing / E. Sousa, I. Graca, T. Baptista [et al.] // Epigenetics. – 2013. – Vol. 8, № 5. – P. 548-558.
- Melo, S.A. Small molecule enoxacin is a cancer-specific growth inhibitor that acts by enhancing TAR RNA-binding protein 2-mediated microRNA processing / S. A. Melo, A. Villanueva, C. Moutinho [et al.] // Proc Natl Acad Sci USA. – 2011. – Vol. 108, № 11. – P. 4394-4399.
- Abell, N. S. Click quantitative mass spectrometry identifies PIWIL3 as a mechanistic target of RNA interference activator enoxacin in cancer cells / N. S. Abell, M. Mercado, T. Caneque [et al.] // J Am Chem Soc. – 2017. – Vol. 139, № 4. – P. 1400-1403.
- Marco, A. Regulatory RNAs in the light of Drosophila genomics / A. Marco // Brief Funct Genomics. – 2012. – Vol. 11, № 5. – P. 356-365.
- Xia, B. Transgenerational programming of longevity and reproduction by post-eclosion dietary manipulation in Drosophila / B. Xia, J.S. de Belle // Aging. – 2016. – Vol. 8, № 5. – P. 1115–1134.
- Hilton, J. F. An algorithm for conducting exact Smirnov tests / J. F. Hilton, C. R. Mehta, N. R. Patel // Computational Statistics & Data Analysis. – 1994. – Vol. 17, № 4. – P. 351–361.
- Martinez, R. L. A pretest for choosing between logrank and wilcoxon tests in the two-sample problem / R. L. M. C. Martinez, J. D. Naranjo // Metron. – 2012. – Vol. 68, № 2. – P. 111–125.
- Wang, C. Statistical methods for testing effects on «maximum lifespan» / C. Wang, Q. Li, D. T. Redden [et al.] // Mech Ageing Dev. – 2004. – Vol. 125, № 9. – P. 629-632.
- Armstrong, R.A. When to use the Bonferroni correction / R.A. Armstrong // Ophthalmic and Physiological Optics. – 2014. – Vol. 34, № 5. – P. 502–508.
- Han, S. K. OASIS 2: online application for survival analysis 2 with features for the analysis of maximal lifespan and healthspan in aging research / S. K. Han, D. Lee, H. Lee [et al.] // Oncotarget. – 2016. – Vol. 7, № 35. – P. 56147-56152.
- Luo, X. Enoxacin inhibits proliferation and invasion of human osteosarcoma cells and reduces bone tumour volume in a murine xenograft model / X. Luo, X. Liu, Q. Tao [et al.] // Oncol Lett. – 2020. – Vol. 20, № 2. – P. 1400-1408.
- Valianatos, G. A small molecule drug promoting miRNA processing induces alternative splicing of MdmX transcript and rescues p53 activity in human cancer cells overexpressing MdmX protein / G. Valianatos, B. Valcikova, K. Growkova [et al.] // PLoS One. – 2017. – Vol. 12, № 10. – P. e0185801.
- Nishi, K. Enoxacin with UVA irradiation induces apoptosis in the AsPC1 human pancreatic cancer cell line through ROS generation / K. Nishi, M. Kato, S. Sakurai [et al.] // Anticancer Res. – 2017. – Vol. 37, № 11. – P. 6211-6214.
- Cao, S. RNA helicase DHX9 may be a therapeutic target in lung cancer and inhibited by enoxacin / S. Cao, R. Sun, W. Wang [et al.] // Am J Transl Res. – 2017. – Vol. 9, № 2. – P. 674-682.
- Ramirez-Moya, J. Impaired microRNA processing by DICER1 downregulation endows thyroid cancer with increased aggressiveness / J. Ramirez-Moya, L. Wert-Lamas, G. Riesco-Eizaguirre [et al.] // Oncogene. – 2019. – Vol. 38, № 27. – P. 5486-5499.
- McDonnell, A. M. Enoxacin and epigallocatechin gallate (EGCG) act synergistically to inhibit the growth of cervical cancer cells in culture / A. M. McDonnell, H. M. Pyles, E. S. Diaz-Cruz [et al.] // Molecules. – 2019. – Vol. 24, № 8. – P. 1580.
- Itoh, A. Enoxacin up-regulates microRNA biogenesis and down-regulates cytotoxic CD8 T-cell function in autoimmune cholangitis / A. Itoh, D. Adams, W. Huang [et al.] // Hepatology. – 2021. – Vol. 74, № 2. – P. 835-846.
- Rocha, A. L. Enoxacin induces oxidative metabolism and mitigates obesity by regulating adipose tissue miRNA expression / A. L. Rocha, T. I. de Lima, G. P. de Souza [et al.] // Sci Adv. – 2020. – Vol. 6, № 49. – P. eabc6250.
- Emde, A. Dysregulated miRNA biogenesis downstream of cellular stress and ALS-causing mutations: a new mechanism for ALS / A. Emde, C. Eitan, L. L. Liou [et al.] // EMBO J. – 2015. – Vol. 34, № 21. – P. :2633-2651.
- McDonnell, A. M. Enoxacin and epigallocatechin gallate (EGCG) act synergistically to inhibit the growth of cervical cancer cells in culture / A. M. McDonnell, H. M. Pyles, E. S. Diaz-Cruz [et al.] // Molecules. – 2019. – Vol. 24, № 8. – P. 1580.
- Xu, Y.P. Zika virus infection induces RNAi-mediated antiviral immunity in human neural progenitors and brain organoids / Y.P. Xu, Y. Qiu, B. Zhang [et al.] // Cell Res. – 2019. – Vol. 29, № 4. – P. 265-273.
- Ahmadi, A. In silico analysis suggests the RNAi-enhancing antibiotic enoxacin as a potential inhibitor of SARSCoV-2 infection / A. Ahmadi, S. Moradi // Sci Rep. – 2021. – Vol. 11, №1. – P. 10271.
- Lyu, B. Enoxacin shows broad-spectrum antiviral activity against diverse viruses by enhancing antiviral RNA interference in insects / B. Lyu, C. Wang, Y. Bie [et al.] // J Virol. – 2022. – Vol. 96, № 4. – P. e0177821.
- Pinto, S. Enoxacin extends lifespan of C. elegans by inhibiting miR-34-5p and promoting mitohormesis / S. Pinto, V. N. Sato, E. A. De-Souza [et al.] // Redox Biol. – 2018. – Vol.18. – P. 84-92.
- Lewis, S.H. Duplication and diversification of Dipteran Argonaute genes, and the evolutionary divergence of piwi and Aubergine / S. H. Lewis, H. Salmela, D. J. Obbard // Genome Biol Evol. – 2016. – Vol. 8, №3. – P. 507-518.
- Malone, C.D. Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary / C.D. Malone, J. Brennecke, M. Dus [et al.] // Cell. – 2009. – Vol. 137, № 3. – P. 522-535.
- Nishida, K.M. Gene silencing mechanisms mediated by Aubergine piRNA complexes in Drosophila male gonad / K. M. Nishida, K. Saito, T. Mori [et al.] // RNA. – 2007. – Vol. 13, № 11. – P. 1911-1922.
- Perera, B. P. U. Somatic expression of piRNA and associated machinery in the mouse identifies short, tissue-specific piRNA / B. P. U. Perera, Z. T. Tsai, M. L. Colwell [et al.] // Epigenetics. – 2019. – Vol. 14, № 5. – P. 504-521.
- Story, B. Defining the expression of piRNA and transposable elements in Drosophila ovarian germline stem cells and somatic support cells / B. Story, X. Ma, K. Ishihara [et al.] // Life Sci Alliance. – 2019. – Vol. 2, № 5. – P. e201800211.


