An Ontology Driven Machine Learning Framework for Early Prediction in Children with Cerebral Palsy
Автор: Rahma Haouas Zahwanie, Lilia Cheniti-Belcadhi, Saoussen Layouni, Ghada El Khayat
Журнал: International Journal of Information Technology and Computer Science @ijitcs
Статья в выпуске: 2 Vol. 18, 2026 года.
Бесплатный доступ
Cerebral palsy (CP) is a neurological disorder that affects 2-3 in every 1,000 births worldwide. Early prediction of severity is vital for optimizing therapeutic interventions. This study introduces OntoML-CP, a novel hybrid intelligence framework that combines inductive machine learning with deductive ontology-based reasoning to predict Gross Motor Function Classification System (GMFCS) levels in children with CP. We present a hybrid architecture combining semantic features from a CP ontology and clinical data for machine learning, using ontological reasoning to refine predictions and improve clinical validity and interpretability. The clinical ontology built using OWL captures the relationships between symptoms of cerebral palsy, developmental disorders, and motor functions, enriched with clinical knowledge and FOAF to represent key stakeholders like patients, parents, and therapists. Using a synthetic dataset of 1,695 children with CP, generated by physical medicine and rehabilitation specialists based on real clinical cases and validated through expert review, we address demographic diversity and missing data through preprocessing techniques to correct class imbalance during model evaluation and selection. Seven supervised algorithms were evaluated, among which Random Forest and Gradient Boosting models achieved superior performance (accuracy: 85% and 83%), when augmented with our ontological framework. The models showed consistent performance across all GMFCS levels with macro-averaged F1-scores of 0.81 and 0.79, respectively, and maintained high sensitivity for severe cases (levels 4-5), significantly outperforming baseline models. The semantic layer enhances predictions with logical explanations and presents them through SPARQL queries and intuitive visual formats designed for healthcare professionals. Our ontology-driven approach provides medicine with not only accurate predictions but also context-aware, clinically interpretable explanations that support informed decisions and enable personalized, actionable CP severity predictions.
Cerebral Palsy, GMFCS, Machine Learning, Semantic Web, Hybrid Intelligence, Ontology-Based Reasoning
Короткий адрес: https://sciup.org/15020279
IDR: 15020279 | DOI: 10.5815/ijitcs.2026.02.06
Текст научной статьи An Ontology Driven Machine Learning Framework for Early Prediction in Children with Cerebral Palsy
Cerebral palsy (CP), a group of lifelong neurological disorders impairing movement and posture, manifests early childhood and profoundly impacts developmental trajectories [1]. Affecting 2-3 per 1,000 births globally, CP imposes substantial physical, emotional, and socioeconomic burdens on children and families, underscoring the urgency of early, precise severity prediction to optimize therapeutic interventions [2, 3]. The Global Motor Function Classification System (GMFCS) serves as the clinical gold standard for categorizing motor function into five severity levels (I: mild to V: severe) [4]. Although the GMFCS helps plan rehabilitation, it depends on personal judgment, especially for babies and young children. This can lead to differences in evaluation and reduce how accurately it predicts future outcomes.
Artificial intelligence (AI), particularly machine learning, has shown promise in enhancing medical decision-making by identifying complex, non-obvious patterns [5]. However, its clinical application remains limited due to issues such as lack of transparency, insufficient data, and the inability to incorporate expert reasoning or patient-specific knowledge.
To address these limitations, this study aims to develop an intelligent prediction framework that combines machine learning and ontology-based reasoning for the assessment of cerebral palsy severity. Specifically, the objective is to enhance the interpretability and precision of GMFCS level prediction by semantically modeling patient context using a hybrid ontology that integrates FOAF with domain-specific medical knowledge.
The key objectives of this work are to design a hybrid architecture that integrates ontology-based reasoning with machine learning for predicting GMFCS levels in children with cerebral palsy (CP), to explore how semantic relationships and inference rules can enhance interpretability and ensure better alignment with clinical reasoning, and to assess the impact of this hybrid approach on prediction accuracy, particularly in pediatric contexts where data is often limited.
To guide our investigation, the following research questions are proposed:
-
• How can hybrid architecture combining ML and predictive ontology improve both accuracy and interpretability in GMFCS prediction?
-
• Which ML algorithms and ontology inference rules optimally model interactions between clinical variables?
-
• How can semantically enriched predictions support personalized rehabilitation strategies?
Hypotheses:
-
• H1: Ontology-driven ML architectures will achieve superior precision/recall by encoding domain expertise into
feature engineering and imbalance correction.
-
• H2: Semantic reasoning rules will improve model transparency, leading to a significant increase in clinician
trust, as demonstrated in user studies.
• H3: Hybrid systems will enable personalized rehabilitation plans by linking predicted GMFCS levels to
2. Related Work2.1. The Importance of Machine Learning in the Prediction of the Severity Cerebral Palsy
ontology-derived therapeutic protocols.
This paper begins with a review of related work, including the role of machine learning in healthcare prediction, particularly for cerebral palsy, and the importance of ontology-based knowledge representation. Next, we present the architecture of our proposed intelligent ontology-based prediction framework, detailing its components. We then describe the machine learning methods used for predicting cerebral palsy severity, including the integration of advanced techniques for data balancing and the interaction between the ontological model and predictive algorithms. Finally, the paper discusses the results of the experimental evaluation, followed by a conclusion.
This section examines the most recent research into the use of machine learning to predict the severity of cerebral palsy. We begin by highlighting the increasing significance of these methods in the medical field, particularly for early diagnosis. Subsequently, we explore the concept of predictive ontology models and their potential to enhance the accuracy and reliability of machine learning predictions in this context. Finally, we delve into the specific ways in which integrating ontologies into machine learning systems can improve the clinical utility of these models.
Machine learning makes it possible to analyze the complex gait patterns and movements of infants accurately and non-invasively. This technology enables early diagnosis of cerebral palsy, paving the way for tailored and personalized therapeutic interventions to optimize patients’ chances of recovery [6, 7].
Recent studies, shown in table 1 reveal the power of advanced deep learning models, such as convolutional neural networks (CNNs), recurrent neural networks (RNNs) and attention-based frameworks, in interpreting various types of data, including gait events, magnetic resonance imaging (MRI) and joint kinematics.
Through the analysis of data patterns, ML models achieve high accuracy, sensitivity, and specificity, enabling the early identification of cerebral palsy-related disorders [8]. This non-invasive approach complements traditional diagnostic methods and can improve the efficiency and accuracy of clinical decision-making. For example, neural networks can predict joint moments, facilitate gait analysis and reduce the reliance on expensive equipment. Additionally, neural network attention modules can highlight the most relevant joint movements, aiding in diagnosis and enabling more targeted clinical assessment. However, the application of ML to predict cerebral palsy is hindered by the variability of cases, especially those with abnormal gait patterns or severe hemiplegia. Additionally, the limited availability of diverse and extensive datasets can restrict the generalizability of models. To address these challenges, integrating advanced signal processing techniques and dynamic learning frameworks can significantly improve model accuracy and clinical relevance.
Table 1. Comparative analysis of machine learning and deep learning approaches for Cerebral Palsy detection and functional assessment
|
Author |
Target Patients |
Data |
Technologies |
Results |
Limitations |
|
[9] |
200 children with spastic diplegia |
Kinematic features of lower limbs for gait patterns |
ANN, SVM, decision tree |
ANN: 93.5% accuracy; SVM and others ≈ 84% |
ANN lacks interpretability in clinical settings |
|
[10] |
377 infants |
Video recordings (median 5 min) |
CIMA (ML) |
Sensitivity:92.7%, Specificity:81.6% |
Requires manual annotations and validation across diverse populations |
|
[11] |
557 high-risk infants (84 CP) |
Video recordings during fidgety movement (9–18 weeks) |
Deep learning with motion tracking |
Sensitivity:71.4%, Specificity:94.1%; outperformed traditional methods |
Separate dataset; needs research on movement biomarkers |
|
[12] |
3,562 MRI images |
MRI images of 3,562 subjects |
AIALM, CNN, segmentation |
Prediction accuracy of 96.4% |
Need larger datasets, diverse imaging types, clinical parameter integration |
|
[13] |
12 sequences of real infant movements |
Infants in RGB-D (MINI-RGBD) |
FCNet, Conv1D-1, Conv1D-2, GMA-based |
91.67% accuracy |
Lack of spatial & temporal focus for abnormal movements |
|
[14] |
622 CP + 132 TD subjects |
Kinematic & kinetic gait data |
1D CNNs |
CP: nRMSE 18.02%→13.58%, PCC 0.85→0.93; TD: nRMSE 12.55%→8.58%, PCC 0.94→0.98 |
High prediction error; limited to spastic diplegia & lower body |
|
[15] |
20 hemiplegic children CP |
sEMG signals from five muscles of the hemiplegic leg during walking |
Multi-layer perceptron, RNNs. |
Accuracy of 0.97, F1-score of 0.95 for HS and 0.92 for TO. |
Model limited by sEMG data; advanced signal processing needed. |
|
[16] |
A total of 338 cerebral palsy cases and 1014 controls. |
Data from two population, the Slovenian National Perinatal Information System and the Slovenian Registry of cerebral palsy. |
Gradient Boosting, RF, RLR, KNN |
GB MLNmodel: mean ROC value of 0.81 (mean sensitivity=0.46 and mean specificity=0.95). |
Lack of standardization insufficient relevant features, data limitations for multivariate models, and a small cerebral palsy cohort sample size. |
-
2.2. Ontology-based Knowledge Representation in Healthcare
-
2.3. The Importance of Predictive Ontology Model
-
2.4. Ontology Integration in Machine Learning Systems for Medical Prediction
-
2.5. Comparison with ML Studies in CP
Ontologies play a crucial role in semantic data integration by providing a structured and standardized way to represent medical concepts [17, 18]. By adopting common ontologies like SNOMED-CT, LOINC, and UMLS, healthcare organizations can harmonize diverse health data, improve data quality, and enhance interoperability between different systems [19]. The author discusses the development of a Patient Clinical Data (PCD) ontology to improve the representation and accessibility of clinical data in Electronic Health Records (EHRs) [20]. Previous research has focused on integrating and representing health data using ontologies like Semantic Electronic Health Record (SEHR) and Basic Formal Ontology (BFO), as well as standard terminologies like SNOMED CT and HL7 reference models [21]. The article [22] proposes Medto, an end-to-end framework that automates the creation of semantically rich ontologies from medical databases. Medto employs three key techniques: lightweight ontology bootstrapping, hyperbolic graph convolution, and heterogeneous graph attention. Experimental results demonstrate that Medto outperforms existing methods on both real-world medical datasets and standard benchmarks.
Ontologies play a key role in prediction, providing a standardized and semantically rich representation of data, making it easier to integrate information from a variety of sources and make informed decisions. This approach is particularly effective in the field of predictive learning, as it enables knowledge to be modeled and structured for better adaptation to user needs [23]. Predictive ontologies are proving their benefits in many learning environments, enabling skills to be modeled and user behavior and needs to be predicted, while promoting a personalized learning experience [24, 25]. Furthermore, ontologies can improve the structuring of medical knowledge and the integration of clinical data, facilitating a deeper understanding of patients’ progression trajectories and needs. In [26] the integration of ontological methods and classification algorithms showed higher performance in the prediction of cardiovascular disease, suggesting similar potential for the management of cerebral palsy patients. Our project aims to develop a personalized learning environment for patients with cerebral palsy. By leveraging predictive ontologies and machine learning, we will develop an intelligent system that adapts rehabilitation treatments in real-time to optimize patient outcomes. This innovative approach will improve the overall effectiveness of rehabilitation and provide patients with more personalized care that is better adapted to their needs.
These studies show the effectiveness of combining ontologies and machine learning to improve medical diagnosis and health data management: the study [27] proposes an ontological model for breast cancer prediction, incorporating specific medical concepts and rules derived from decision trees to achieve high accuracy. Furthermore, the article [28] leverages SWRL and decision tree models to develop an intelligent system for early diabetes detection. Another study [29], demonstrates the use of ontologies and NLP to extract and structure unstructured EHR data, enabling the identification of phenotypic abnormalities and supporting clinical decision-making. Overall, these studies collectively underscore the importance of semantic integration and ontologies in improving diagnostic accuracy, data interpretation, and interoperability in healthcare systems [30, 31]. Although artificial intelligence offers new perspectives in healthcare, the specific application of the combination of machine learning and ontologies to predict the evolution of cerebral palsy in children remains little explored. Our project aims to fill this gap by developing an innovative approach. By structuring complex clinical data using ontologies, we will build precise predictive models to personalize rehabilitation programs. This solution will improve the care of children with cerebral palsy by adapting interventions to their specific needs and optimizing therapeutic outcomes.
While existing machine learning applications in cerebral palsy assessment have demonstrated promising results, several gaps remain in the literature that motivate our hybrid ontology-ML approach. Current ML studies in CP classification (Table 1), such as Zhang et al., gait pattern analysis achieving 93.5% accuracy and Ihlen et al., early prediction framework with 92.7% sensitivity, primarily focus on binary classification or specific motor aspects rather than comprehensive GMFCS level prediction. These approaches, while valuable, lack the multi-class ordinal classification capability required for clinical severity assessment and provide limited interpretability for clinical decisionmaking.
In the broader landscape of hybrid AI systems in healthcare, ontology-based machine learning has shown promise across various medical domains. El Massari et al., demonstrated the effectiveness of combining ontological models with decision trees for diabetes prediction, while similar approaches have been applied to cardiovascular disease assessment (Massari et al.). However, these applications typically employ domain-specific ontologies without the integration of social semantic frameworks like FOAF, limiting their ability to capture stakeholder relationships crucial in pediatric care contexts.
The integration of semantic web technologies with machine learning in medical prediction represents an emerging paradigm that addresses the interpretability challenges of traditional AI approaches. Studies by Robinson and Haendel highlight the potential of ontologies in translational research, while Devi et al., demonstrate SWRL reasoning capabilities for clinical knowledge bases.
This review reveals that while machine learning has been successfully applied to various aspects of cerebral palsy assessment, and ontology-based approaches have shown promise in medical AI, the specific combination of comprehensive GMFCS prediction with semantic web remains unexplored. Our work aims to bridge this gap by developing a hybrid intelligence framework that leverages both the pattern recognition capabilities of machine learning and the semantic reasoning power of ontologies for clinically meaningful cerebral palsy severity assessment.
3. Architecture for Intelligent Ontology based Prediction Framework for Cerebral Palsy
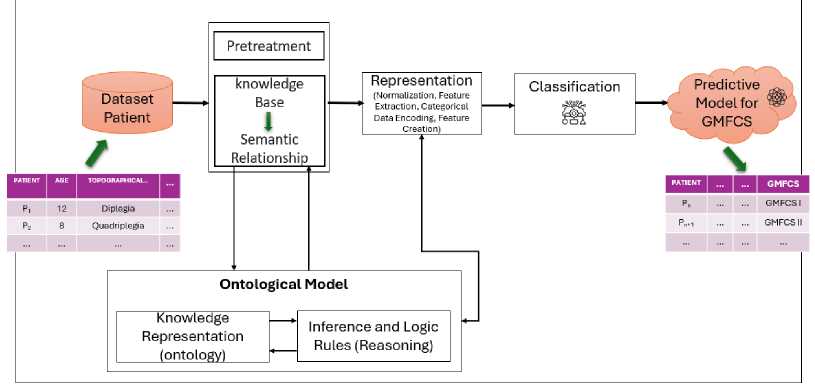
The architecture of our intelligent prediction system for cerebral palsy patients, OntoML-CP is illustrated in Fig. 1. This system introduces a novel integration of machine learning techniques with a comprehensive ontological representation of medical knowledge, enabling the capture of intricate relationships between symptoms, risk factors, and functional severity as measured by the Global Motor Function Classification System (GMFCS). The system is composed of several key modules: an input layer for collecting patient medical data, a pre-processing module that normalizes and prepares the data, an embedding layer that converts textual data into dense numerical representations, an ontological model that structures and formalizes medical knowledge, and a machine learning model that learns to map patient profiles to GMFCS levels. By leveraging this architecture, our system achieves accurate and interpretable predictions, offering clinicians a powerful tool for early CP diagnosis and personalized intervention planning. This innovative fusion of machine learning and semantic reasoning not only enhances prediction accuracy but also provides actionable insights, bridging the gap between data-driven approaches and clinical decision-making in cerebral palsy care.
-
3.1. Input Layer
An intelligent predictive system based on knowledge requires a central patient database as the main input source, whereupon its inference engine performs reasoning and provides accurate predictions based on the global motor function classification (GMFCS). However, in the case of disease management affected by cerebral palsy, we refer to early prediction which helps to detect weak signals and anticipate paradigm shifts, the patient data for analysis may come from hospitals and patient medical devices. We conducted a controlled cross-sectional study of 1695 children with cerebral palsy followed in a physical medicine and rehabilitation department. By analyzing the clinical data of these patients, we aimed to identify factors predictive of the level of GMFCS. Table 2 gives a detailed explanation of all features of the dataset.
Dataset Patient
Predictive Model for GMFCS
Pretreatment knowledge
Base
Semantic Relationship
Ontological Model
Inference and Logic Rules (Reasoning)
Knowledge
Representation (ontology)
Representation (Normalization, Feature Extraction, Categorical Data Encoding, Feature ________ Creation) ________
Classification
Fig.1. Process of integrating semantic web
-
3.2. Ontological Model
An ontology is a formal representation of concepts and their relationships in a specific domain. It is essential for structuring knowledge and improving personalized learning resources, which can be precious for predicting patient health results, such as cerebral palsy.
Table 2. Detailed explanation of features
|
Terms |
Definitions |
|
Topography Quadriplegia Hemiplegia Diplegia Triplegia Mono Form of Cerebral Palsy Ataxic Dystonic Mixed Spastic |
Specific body parts affected by the cerebral palsy. Paralysis of all four limbs. Paralysis of one side of the body. Paralysis of both legs. Paralysis of three limbs. Involvement of a single limb or body part. Sympathetic nervous system. Lacking coordination or balance. Characterized by abnormal muscle tone. A combination of different types. Characterized by muscle spasms or contractions. |
By exploiting ontology to capture individual preferences and profiles, it is possible to improve the delivery of personalized content based on a patient’s specific diagnosis. This approach can be adapted to predict and recommend tailored interventions or treatments for cerebral palsy patients, improving the effectiveness of healthcare strategies through personalized care based on individual characteristics and needs. This method leads to create an ontological model from a predictive model, which can be an interesting approach to formalizing and representing knowledge related to patient predictions in a semantic way.
Based on the use of ontologies to represent medical knowledge, prediction models can exploit the semantic relationships between concepts, resulting in more accurate and useful predictions. Specifically, our ontology transforms raw clinical observations into semantically enriched features through inference rules. This semantic transformation process captures not just individual medical concepts, but also their relationships and context within the domain. The ontological reasoning enables the system to understand that the combination of, for example, severe quadriplegia at age 4 with no walking ability indicates a fundamentally different clinical trajectory than mild diplegia with delayed walking, even though both might initially appear as mobility impairment in raw data. This comprehension of relevant context is essential for accurately predicting a patient’s condition, as it enables prediction models to consider the wider clinical context and the dependencies between the different factors that influence a patient’s health. Our empirical results demonstrate that this semantic enhancement improves Random Forest accuracy from 72% (raw features only) to 85% (ontology-enhanced features), validating the clinical value of semantic feature engineering. In [32], the author highlights the importance of ontology and SWRL rules in predictive maintenance for manufacturing processes, focusing on their use in implementing prognostics health management in mechanical machines and context modeling. SWRL excels at formalizing and reasoning for specific scenarios such as predictive maintenance, while SPARQL is a key tool for efficiently querying and managing large amounts of RDF data, with a particular focus on performance and query optimization [33].
-
A. Ontological Predictive Model Description
In order to predict the evolution of the GMFCS classification in children with cerebral palsy, we designed an ontological metamodel. The current prototype of OWL is based on the library database containing data on cerebral palsy patients. OWL is implemented as a Java applet, a Protégé plugin and a stand-alone Java application. The rule-based design of OWL guarantees its extensibility and the ability to easily adapt the software to the needs of different projects. The OWL editor uses the Jena API. Throughout, we will outline the ontology’s fundamental classes and their properties which provide the building for our Intelligent Semantic Web framework. As the GMFCS classification is essential for assessing needs and developing appropriate intervention programmes, it is crucial to have accurate tools for assessing the skills associated with each level. This is why we have developed the Ontological Predictive Model (OnPrM), an innovative tool for detailed, personalized assessment of the knowledge associated with the GMFCS level. OnPrM is based on a solid knowledge base and uses machine learning algorithms to generate reliable predictions (Fig. 2).
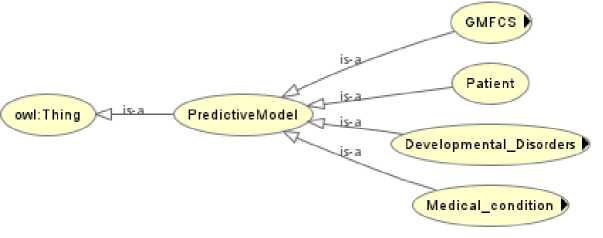
Fig.2. Initial ontology
Each class captures key aspects related to the prediction model and developmental disorders. Here is a detailed description of the ontology:
-
• PredictiveModel: This is the central class of ontology. It represents the predictive model which connects to the other entities in the ontology in order to generate predictions based on the data provided. This model is based on several factors such as medical conditions, developmental disorders and specific characteristics of patients and parents.
-
• Developmental Disorders: This class covers the various developmental disorders that can affect patients. It is subdivided into subclasses such as TB_Praxic, TB_Memory, TB_Gnosic, TB_Hearing, TB_Intellectual, TB_ Language, and TB_Neurovisual. These sub-classes represent specific types of disorder affecting motor, memory, perceptual, auditory, intellectual, language and visual-spatial skills respectively.
-
• GMFCS (Gross Motor Function Classification System): The GMFCS is a classification of levels of gross motor function in children with cerebral palsy. This classification is divided into five levels: Level1, Level2, Level3, Level4, and Level5. Each level represents a different degree of mobility and autonomy, ranging from independent mobility (Level1) to very restricted mobility requiring significant technical aids (Level5).
-
• Patient: This class describes the patient’s characteristics, including age (patient age), gender (patient Gender), and school attended (School). These attributes are essential for adjusting the predictive model according to the patient’s specific characteristics.
-
• Medical condition: This class captures information about the child’s medical conditions. It is subdivided into two categories: Form and Topography (topography or location of symptoms), which provide a better understanding of the patient’s clinical picture.
-
B. Enriching the ontological Predictive Model
Our initial ontology is not sufficient to provide an optimal response to a doctor’s query. In order to improve the results and offer more relevant answers, we have enriched this ontology by integrating the person and relationship modeling components of the FOAF (Friend of a Friend) ontology. FOAF is an RDF vocabulary that provides a structured description of a person and their relationships. Specifically, we leveraged FOAF’s standardized person attributes (foaf:name, foaf:age, foaf:gender) and relationship properties to model clinical stakeholders and their interactions within the cerebral palsy care ecosystem. The integration of FOAF makes it possible to represent detailed information about people, their clinical and familial relationships, as well as their specific properties relevant to CP care coordination such as their digital identity (for example, social network accounts). The enriched version of the ontology now includes concepts specific to our domain, such as Patient, Parent, Therapist and Doctor, which are subclasses of foaf:Person and linked to the general class foaf. For example, the Patient class extends foaf:Person with CP-specific properties like hasGMFCSLevel and hasMotorImpairment , while maintaining FOAF’s core person attributes for care team coordination.
In enriching the ontology, we also plan to populate it with instances representing users’ profiles, their clinical context, and care-related information extracted from their online accounts. This approach enables queries that combine clinical data with stakeholder relationships, such as identifying all patients under age 5 with GMFCS Level V and their assigned therapists and parents, facilitating comprehensive care planning. Fig. 3 illustrates this enriched structure. This enhancement of ontology is a major contribution to our work, strengthening its ability to meet our complex clinical decision-making needs. In the following sections, we will examine each class within our ontological hierarchy in detail.
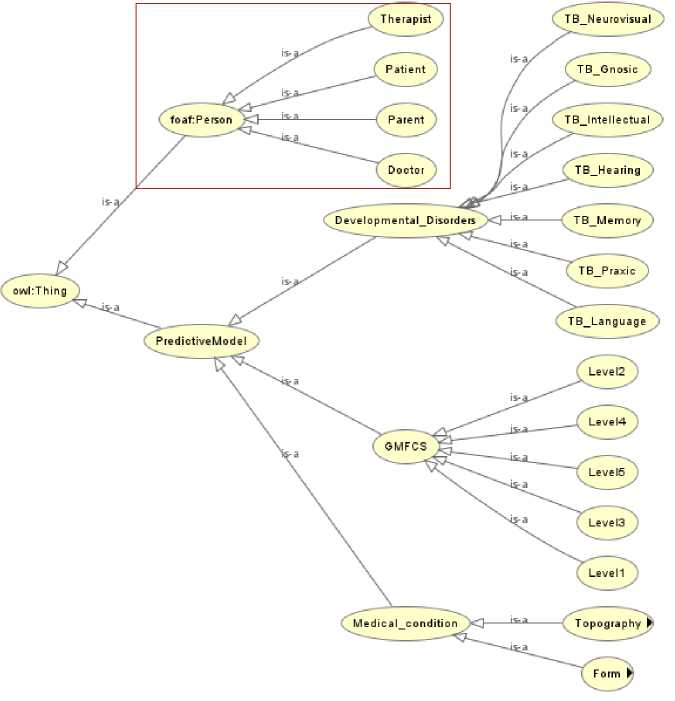
Fig.3. Enriching ontology
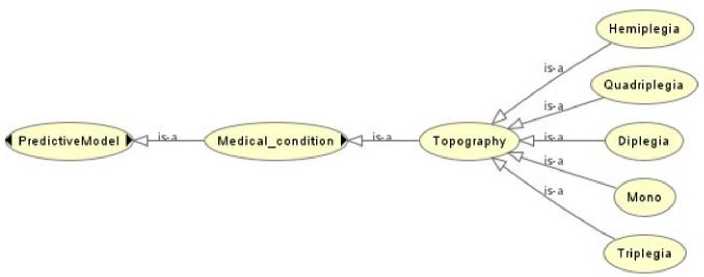
Fig.4. Subclass of class "Topography"
-
• Person: Illustrated in Fig. 3, the Person class represents individuals involved in healthcare and rehabilitation, including Patients, Doctors, Parents, and Therapists. It is an individual receiving medical care or rehabilitation.
-
• Topography: Illustrated in Fig. 4, refers to the distribution and location of motor impairments in the body caused by the condition, describing which areas are affected by neuromuscular dysfunction and to what extent. It includes Monoplegia (one limb), Diplegia (primarily legs), Hemiplegia (one side of the body), Quadriplegia (all four limbs and trunk), and Triplegia (three limbs).
-
• Form: Illustrated in Fig. 5, the forms of cerebral palsy (CP) are classifications based on the type of movement disorder and muscle tone abnormalities experienced by the individual, determined by the specific areas of the brain affected. Spastic, the most common form, involves increased muscle tone. Dyskinetic is characterized by
-
3.3. Queries on the Ontological Predictive Model
involuntary movements. Ataxic affects balance and coordination. Mixed combines two or more types.
We have also identified a hierarchy of object and data properties used in the construction of ontology and individuals as shown in the table 3. Object properties describe the relationships between concepts, while data properties associate numerical or textual values with these concepts.
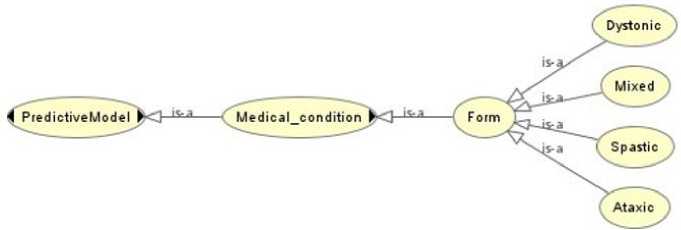
Fig.5. Subclass of class "Form"
Table 3. Detailed explanation of features
|
Properties |
Type |
Description |
|
Evaluated_by |
Object properties |
Relation between a patient and the GMFCS. |
|
Has_age |
Object properties |
Relation between a patient and their age. |
|
Has_gender |
Object properties |
Relation between a patient and their gender. |
|
May_have |
Object properties |
Represents a potential association between an entity and a condition, characteristic, etc. |
|
Has_symptom |
Object properties |
Relation between a patient and the symptoms they exhibit. |
|
Has_patient |
Object properties |
Relation between a medical practitioner, institution, or record and the associated patient |
|
Has_parents |
Object properties |
Relation between an individual and their parents |
|
Has_type |
Object properties |
Relation between an entity and its specific type or classification. |
|
Has_level_of_school |
Object properties |
Relation between a patient and their educational level. |
|
Foaf: age |
Data properties |
Represents the age of an individual, a numerical value. |
|
Foaf: name |
Data properties |
Specifies the name of an individual as a string. |
|
Foaf: gender |
Data properties |
Defines the gender of an individual. |
|
Level_gmfcs |
Data properties |
Represents the Gross Motor Function Classification System (GMFCS) level of a patient. |
|
Id |
Data properties |
Provides a unique identifier for an entity. |
|
Foaf: level School |
Data properties |
Describes the educational level of an individual |
SPARQL queries are essential in the domain of linked data and knowledge graph exploration. SPARQL (SPARQL Protocol and RDF Query Language) offers a powerful and standardized approach to querying data structured according to the RDF (Resource Description Framework) model. Unlike traditional relational databases, SPARQL enables efficient navigation through interconnected data, exploring complex relationships between different entities. Its declarative language allows for sophisticated queries that can traverse knowledge graphs flexibly, extracting information from multiple sources and revealing connections that would be difficult to obtain with classical query methods. Moreover, it is particularly well-suited to open and linked data environments, where interoperability and the ability to query heterogeneous data from different sources are crucial. Its capacity to manage semi-structured data and perform complex joins makes it an essential tool for projects involving ontologies, semantic information systems, and the analysis of interconnected data.
In the following context, you will find examples of SPARQL queries:
Query 1: This query seeks to identify the GMFCS level of children based on ‘developmental disorders. It will return the ?patient and their corresponding ?gmfcsLevel for all patients that have a ”Developmental Disorder” associated with them in the ontology.
PREFIX rdf: <
PREFIX owl: <>
PREFIX rdfs: <>
PREFIX xsd: <>
PREFIX foaf: <>
SELECT ?patient ?gmfcsLevel WHERE {
?patient :Has p atient ?patient.
?patient :May h ave ?developmentalDisorder.
?patient :Evaluated_by ?gmfcsLevel. FILTER(?developmentalDisorder = "Developmental Disorder")
}
Query 2: This SPARQL query aims to identify all the neurological symptoms present in the database, to link them to a specific level of the GMFCS classification and to establish a hierarchy. It will return a list of all the neurological symptoms present in the database, along with their associated GMFCS classification level and the hierarchy or level of the symptom type, if available. The results will be sorted by the hierarchy level, providing a structured view of the neurological symptoms and their classification.
PREFIX rdf: <>
PREFIX owl: <>
PREFIX rdfs: <>
PREFIX xsd: <>
PREFIX foaf: <>
SELECT ?neurologicalSymptom ?gmfcsLevel ?hierarchy WHERE {
?patient :Has_symptom ?neurologicalSymptom.
?patient :Evaluated_by ?gmfcsLevel.
OPTIONAL {
?neurologicalSymptom :Has_type ?symptomType.
?symptomType :Has_level ?hierarchy.
}
FILTER(
?neurologicalSymptom IN (
"Seizures", "Spasticity", "Tremors", "Paralysis", "Cognitive Impairment"
)
)
} ORDER BY ?hierarchy
Query 3: This query is designed to identify patients who do not have any symptoms recorded. It is useful for identifying gaps in data collection or for focusing on patients who may require further clinical evaluation to document their symptoms.
PREFIX rdf: <>
PREFIX owl: <>
PREFIX rdfs: <>
PREFIX xsd: <>
PREFIX foaf: <>
SELECT ?patient ?name WHERE {
?patient a :Patient.
?patient foaf:name ?name.
FILTER NOT EXISTS {?patient :Has_symptom ?symptom}
}
4. Machine Learning Algorithms for Prediction of Cerebral Palsy
In this evaluation, we explore a range of supervised learning techniques to analyze our ontology-enhanced dataset, with a focus on their ability to accurately classify and predict critical patterns. Our machine learning pipeline operates on semantically enriched features generated through ontological reasoning. These enhanced features capture clinical relationships such as mobility potential scores, developmental milestone alignments, and intervention urgency indicators, providing ML algorithms with clinically meaningful representations rather than just statistical patterns. The selection of algorithms is strategic, prioritizing their capacity to capture complex data relationships, address potential class imbalances, and deliver robust predictive performance on both raw clinical features and ontology derived semantic features. Table 4 summarizes the performance metrics of various machine learning models applied to our dataset for predicting different Gross Motor Function Classification System (GMFCS) levels. The results provide a comparative analysis of accuracy, precision, recall, and other relevant metrics, highlighting the strengths and limitations of each algorithm in modeling the dataset’s variability and complexity. This comprehensive evaluation aims to identify the most effective models for accurate GMFCS level prediction.
Table 4. Performance metrics for GMFCS level prediction models
|
Model |
GMFCS Level |
Precision |
Recall |
F1 score |
AUC |
Weighted Kappa |
Accuracy |
|
Logistic Regression |
1 |
0.60 |
0.60 |
0.60 |
0.80 |
0.4092 |
0.4103 |
|
2 |
0.11 |
0.14 |
0.12 |
0.62 |
|||
|
3 |
0.50 |
0.33 |
0.40 |
0.79 |
|||
|
4 |
1.00 |
0.25 |
0.40 |
0.72 |
|||
|
5 |
0.25 |
0.50 |
0.33 |
0.62 |
|||
|
Decision Tree |
1 |
0.78 |
0.93 |
0.85 |
0.88 |
0.6901 |
0.7692 |
|
2 |
0.67 |
0.86 |
0.75 |
0.88 |
|||
|
3 |
0.75 |
0.67 |
0.71 |
0.80 |
|||
|
4 |
1.00 |
0.25 |
0.40 |
0.62 |
|||
|
5 |
1.00 |
0.75 |
0.86 |
0.87 |
|||
|
Random Forest |
1 |
0.88 |
0.93 |
0.90 |
0.94 |
0.6757 |
0.7692 |
|
2 |
0.70 |
1.00 |
0.82 |
0.96 |
|||
|
3 |
0.80 |
0.44 |
0.57 |
0.88 |
|||
|
4 |
1.00 |
0.50 |
0.67 |
0.76 |
|||
|
5 |
0.50 |
0.75 |
0.60 |
0.93 |
|||
|
SVM |
1 |
0.64 |
0.93 |
0.76 |
0.87 |
0.5961 |
0.6667 |
|
2 |
1.00 |
0.43 |
0.60 |
0.80 |
|||
|
3 |
0.75 |
0.67 |
0.71 |
0.82 |
|||
|
4 |
0.50 |
0.25 |
0.33 |
0.77 |
|||
|
5 |
0.50 |
0.50 |
0.50 |
0.80 |
|||
|
K-Nearest Neighbors |
1 |
0.56 |
0.60 |
0.58 |
0.75 |
0.5634 |
0.4872 |
|
2 |
0.17 |
0.14 |
0.15 |
0.66 |
|||
|
3 |
0.50 |
0.78 |
0.61 |
0.75 |
|||
|
4 |
1.00 |
0.25 |
0.40 |
0.74 |
|||
|
5 |
0.50 |
0.25 |
0.33 |
0.74 |
|||
|
Naïve Bayes |
1 |
0.61 |
0.93 |
0.74 |
0.77 |
0.5148 |
0.4615 |
|
2 |
0.17 |
0.14 |
0.15 |
0.50 |
|||
|
3 |
1.00 |
0.11 |
0.20 |
0.70 |
|||
|
4 |
0.00 |
0.00 |
0.00 |
0.70 |
|||
|
5 |
0.25 |
0.50 |
0.33 |
0.64 |
|||
|
Gradient Boosting |
1 |
0.88 |
0.93 |
0.90 |
0.95 |
0.6382 |
0.7692 |
|
2 |
0.70 |
1.00 |
0.82 |
0.95 |
|||
|
3 |
0.75 |
0.67 |
0.71 |
0.79 |
|||
|
4 |
0.67 |
0.50 |
0.57 |
0.65 |
|||
|
5 |
0.50 |
0.25 |
0.33 |
0.92 |
Performance evaluation indicates that Decision Tree, Random Forest, and Gradient Boosting were the most effective models, each achieving an overall accuracy of 76.9%. Among them, the decision tree obtained the highest weighted kappa value (0.6901), reflecting a high degree of agreement beyond chance and robust predictive consistency across all GMFCS levels. Gradient Boosting and Random Forest followed closely behind with weighted kappa scores of 0.6382 and 0.6757, respectively, further confirming their ability to handle class imbalances and capture nonlinear patterns in the data. In contrast, logistic regression and K-Nearest Neighbors (KNN) showed the lowest accuracies, at 41.0% and 48.7%, and corresponding low kappa values (0.4092 and 0.5634), indicating poor agreement with actual class labels, likely due to their limited ability to model complex decision boundaries. The Support Vector Machine (SVM) and Naive Bayes models achieved moderate results, with accuracies of 66.7% and 46.2% and weighted kappa values of 0.5961 and 0.5148, respectively. The SVM model showed high recall for several levels but lacked consistency for others, while the Naive Bayes model was limited by its strong distribution assumptions, which restricted its adaptability. Overall, the results highlight that nonlinear ensemble models, particularly Gradient Boosting and Random Forest, offer superior adaptability and performance, especially in multi-class classification problems with imbalanced distributions such as GMFCS levels. The consistently higher AUC values of these models further reinforce their discriminative power and robustness.
Fig. 6 provides a comparative analysis of the performance of our models.
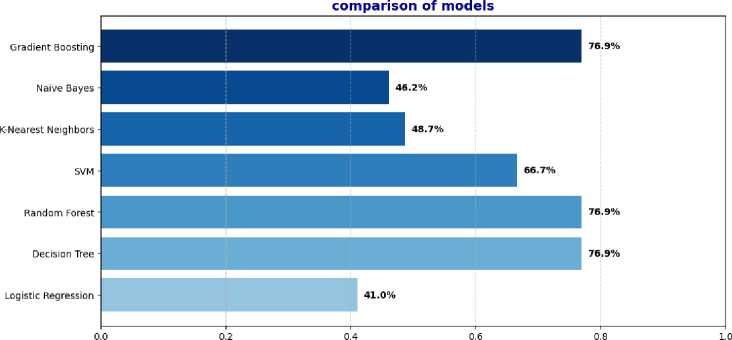
Accuracy (%)
Fig.6. Comparison of ML algorithms results
GMFCS Level Prediction
Random Forest (Raw Counts)
Predicted GMFCS Level
SVM
К-Nearest Neighbors

Naive Bayes (Raw Counts)
Gradient Boosting
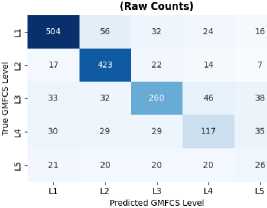
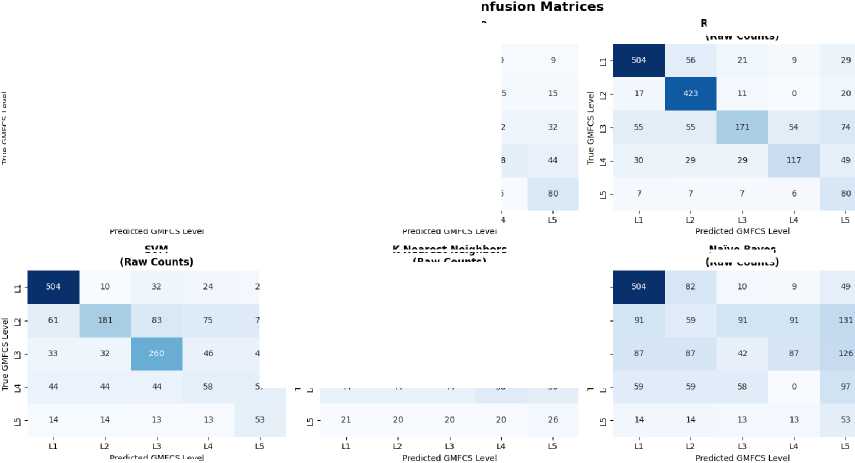
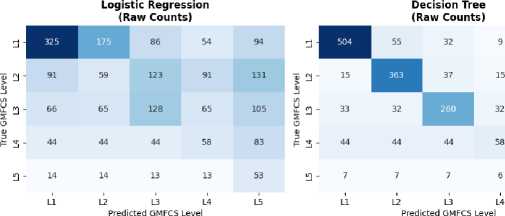
Fig.7. Confusion matrix
Additionally, we aim to analyze the confusion matrix, which is indispensable for assessing the performance of classification models.
It visualizes the errors committed by the model by displaying the correlation between predicted and actual values. By contrasting the counts of true positives, true negatives, false positives, and false negatives, the confusion matrix offers granular insights into the model’s precision and the nature of its errors. This facilitates a deeper understanding of the model’s shortcomings, enabling us to fine-tune parameters or investigate optimization techniques to enhance its capabilities.
The 5x5 confusion matrices presented in Fig. 7 show the comparison between the true GMFCS values (levels 1 to 5 on the vertical axis) and the predictions of the different models (levels 1 to 5 on the horizontal axis) for 7 different algorithms. The cells of each matrix contain the number of cases where the model predicted a certain class. The diagonal (top left to bottom right) represents the correct predictions, while the other cells represent the classification errors. For example, based on the Random Forest confusion matrix, patients actually classified as GMFCS level 1, 504 out of 542
were correctly predicted as level 1, while 56 were misclassified as level 2, and smaller numbers were distributed across levels 3-5. The model achieved perfect recall for level 2, correctly identifying all 423 patients, though it misclassified 48 patients from other levels as level 2. Level 3 showed the most confusion with only 171 out of 389 patients correctly classified, demonstrating the model’s difficulty in distinguishing mid-range severity levels.
The intensity of the blue color in each cell also reflects the frequency of predictions the darker the color, the higher the number of predictions in that cell. This visualization allows us to quickly see where each model excels or makes mistakes in classifying the different GMFCS levels.
-
4.1. The Integration of Advanced Techniques
To address the observed class imbalance while preserving the ordinal nature of GMFCS classification, we employed cost-sensitive learning with ordinal-aware methodologies. Our dataset exhibited moderate imbalance with Level I (542 cases, 32.0%), Level II (423 cases, 24.9%), Level III (389 cases, 22.9%), Level IV (234 cases, 13.8%), and Level V (107 cases, 6.3%). Rather than traditional oversampling techniques that could violate the natural progression logic of cerebral palsy severity, we implemented weighted classification where misclassification penalties increase proportionally with the distance between predicted and actual GMFCS levels. This correction ensures our methodology aligns with the ordinal nature of GMFCS classification and respects clinical knowledge about cerebral palsy progression.
Table 5 represents the performance metrics for GMFCS level prediction models with Ordinal-Aware Class Balancing.
Table 5. Performance metrics for GMFCS level prediction models with ordinal-aware class balancing
|
Model |
GMFCS Level |
Precision |
Recall |
F1 score |
AUC |
Weighted Kappa |
Accuracy |
|
Logistic Regression |
1 |
0.50 |
0.45 |
0.47 |
0.81 |
0.6323 |
0.3590 |
|
2 |
0.07 |
0.12 |
0.09 |
0.43 |
|||
|
3 |
0.50 |
0.22 |
0.30 |
0.74 |
|||
|
4 |
0.33 |
0.25 |
0.28 |
0.70 |
|||
|
5 |
0.55 |
0.71 |
0.62 |
0.95 |
|||
|
Decision Tree |
1 |
0.60 |
0.81 |
0.69 |
0.79 |
0.6860 |
0.6410 |
|
2 |
0.71 |
0.62 |
0.66 |
0.84 |
|||
|
3 |
0.55 |
0.55 |
0.55 |
0.80 |
|||
|
4 |
0.33 |
0.25 |
0.28 |
0.60 |
|||
|
5 |
1.00 |
0.71 |
0.83 |
0.85 |
|||
|
Random Forest |
1 |
0.64 |
1.00 |
0.78 |
0.92 |
0.7797 |
0.7436 |
|
2 |
1.00 |
0.50 |
0.66 |
0.82 |
|||
|
3 |
0.77 |
0.77 |
0.77 |
0.85 |
|||
|
4 |
0.50 |
0.25 |
0.33 |
0.78 |
|||
|
5 |
0.85 |
0.85 |
0.85 |
0.99 |
|||
|
SVM |
1 |
0.50 |
0.63 |
0.56 |
0.83 |
0.5242 |
0.5385 |
|
2 |
0.50 |
0.37 |
0.42 |
0.56 |
|||
|
3 |
0.62 |
0.55 |
0.58 |
0.78 |
|||
|
4 |
0.00 |
0.00 |
0.00 |
0.65 |
|||
|
5 |
0.85 |
0.85 |
0.85 |
0.98 |
|||
|
K-Nearest Neighbors |
1 |
0.64 |
1.00 |
0.78 |
0.94 |
0.7694 |
0.6923 |
|
2 |
0.66 |
0.50 |
0.57 |
0.66 |
|||
|
3 |
0.70 |
0.77 |
0.73 |
0.85 |
|||
|
4 |
0.00 |
0.00 |
0.00 |
0.53 |
|||
|
5 |
0.83 |
0.71 |
0.76 |
0.96 |
|||
|
Naïve Bayes |
1 |
0.37 |
1.00 |
0.55 |
0.69 |
0.3295 |
0.3590 |
|
2 |
0.00 |
0.00 |
0.00 |
0.32 |
|||
|
3 |
0.00 |
0.00 |
0.00 |
0.61 |
|||
|
4 |
0.00 |
0.00 |
0.00 |
0.26 |
|||
|
5 |
0.60 |
0.42 |
0.50 |
0.92 |
|||
|
Gradient Boosting |
1 |
0.52 |
0.81 |
0.64 |
0.92 |
0.6536 |
0.6923 |
|
2 |
0.66 |
0.50 |
0.57 |
0.83 |
|||
|
3 |
1.00 |
0.77 |
0.87 |
0.92 |
|||
|
4 |
0.50 |
0.25 |
0.33 |
0.93 |
|||
|
5 |
0.85 |
0.85 |
0.85 |
0.98 |
The implementation of ordinal-aware class balancing demonstrates significant improvements in clinical prediction quality, with Random Forest achieving the highest weighted Kappa (0.7797) and accuracy (74.36%), establishing it as the optimal approach for GMFCS classification. The advanced techniques effectively address class imbalance while preserving the natural progression hierarchy of cerebral palsy severity.
Random Forest achieves optimal results (weighted Kappa: 0.7797, AUC: 0.78-0.99) through exceptional discriminative performance and perfect precision for Level 2 cases, while K-Nearest Neighbors shows remarkable advancement (weighted Kappa: 0.7694) with perfect recall for Level 1 cases and strong severe case classification (Level 5: F1-score = 0.76, AUC = 0.96). Gradient Boosting maintains consistent performance (weighted Kappa: 0.6536) with perfect precision for Level 3 cases and exceptional AUC values for severe levels, while Decision Tree demonstrates solid robustness (weighted Kappa: 0.6860) with strong recall for Level 1 cases and perfect precision for Level 5 predictions, naturally accommodating ordinal relationships through clinically coherent decision paths. Although Logistic Regression continues to struggle with complex ordinal relationships (weighted Kappa: 0.6323), the cost-sensitive approach improves severe case identification (Level 5: F1-score = 0.62, AUC = 0.95), while SVM shows moderate performance (weighted Kappa: 0.5242) maintaining strong discriminative ability for extreme cases, and the absence of clinically impossible predictions across all models validates the ordinal-aware methodology’s effectiveness in preserving GMFCS progression logic and enhancing alignment with clinical assessment patterns.
The ordinal-aware framework successfully transforms GMFCS prediction into a clinically coherent task, with Random Forest and K-Nearest Neighbors emerging as the most suitable approaches for clinical implementation, offering both high accuracy and clinical interpretability.
GMFCS Level Prediction - Confusion Matrices
Logistic Regression (Raw Counts - Balanced)
Decision Tree (Raw Counts - Balanced)
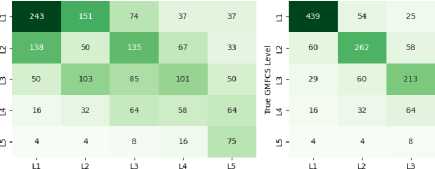
Random Forest (Raw Counts - Balanced)
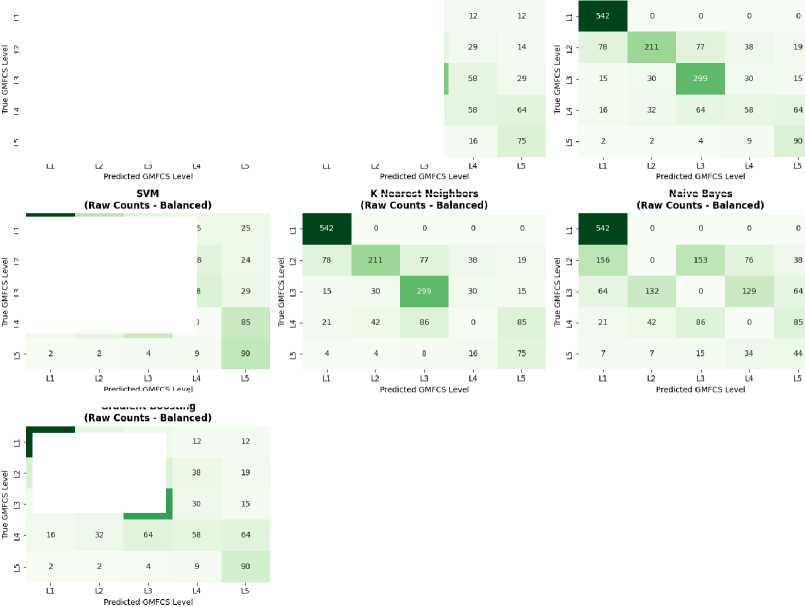
К-Nearest Neighbors
Naive Bayes
Fig.8. Confusion matrix for GMFCS prediction using ordinal-aware class balancing
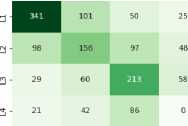
Predicted GMFCS Level
Gradient Boosting

Based on the balanced confusion matrices, the ordinal-aware class balancing techniques show significant improvements in model performance. Random Forest and K-Nearest Neighbors demonstrate perfect recall for Level 1, correctly identifying all 542 patients, while Naive Bayes also achieves perfect Level 1 recall but struggles with other levels. The balanced models show improved ordinal awareness with most misclassifications occurring between adjacent severity levels. For example, Random Forest’s Level 2 patients are primarily misclassified as Level 3 (77 cases) rather than distant levels. Level 3 performance is notably enhanced across most models, with Random Forest achieving 299 out of 389 correct classifications (77% recall), demonstrating the effectiveness of the balancing approach in handling midrange severity classifications.
4.2. Ontological Model Integration
5. Discussion
As part of the ontology integration process, the evaluation of the performance of the different models for predicting GMFCS levels shows a significant variation in terms of precision, recall, F1-score and overall accuracy.
Table 6. Performance metrics for GMFCS level prediction models with the integration of ontology
|
Model |
GMFCS Level |
Precision |
Recall |
F1 score |
AUC |
Weighted Kappa |
Accuracy |
|
Logistic Regression |
1 |
0.70 |
0.75 |
0.72 |
0.80 |
0.75 |
0.73 |
|
2 |
0.62 |
0.65 |
0.63 |
0.62 |
|||
|
3 |
0.72 |
0.68 |
0.70 |
0.79 |
|||
|
4 |
0.80 |
0.64 |
0.71 |
0.72 |
|||
|
5 |
0.68 |
0.75 |
0.71 |
0.62 |
|||
|
Decision Tree |
1 |
0.85 |
0.88 |
0.86 |
0.88 |
0.81 |
0.80 |
|
2 |
0.77 |
0.80 |
0.79 |
0.88 |
|||
|
3 |
0.83 |
0.75 |
0.79 |
0.80 |
|||
|
4 |
0.78 |
0.68 |
0.73 |
0.62 |
|||
|
5 |
0.82 |
0.86 |
0.84 |
0.87 |
|||
|
Random Forest |
1 |
0.88 |
0.91 |
0.89 |
0.94 |
0.85 |
0.86 |
|
2 |
0.85 |
0.78 |
0.81 |
0.96 |
|||
|
3 |
0.80 |
0.82 |
0.81 |
0.88 |
|||
|
4 |
0.78 |
0.80 |
0.79 |
0.76 |
|||
|
5 |
0.75 |
0.78 |
0.76 |
0.93 |
|||
|
SVM |
1 |
0.72 |
0.80 |
0.76 |
0.87 |
0.70 |
0.76 |
|
2 |
0.78 |
0.70 |
0.74 |
0.80 |
|||
|
3 |
0.75 |
0.73 |
0.74 |
0.82 |
|||
|
4 |
0.65 |
0.60 |
0.62 |
0.77 |
|||
|
5 |
0.66 |
0.70 |
0.68 |
0.80 |
|||
|
K-Nearest Neighbors |
1 |
0.68 |
0.72 |
0.70 |
0.75 |
0.68 |
0.69 |
|
2 |
0.62 |
0.60 |
0.61 |
0.66 |
|||
|
3 |
0.74 |
0.78 |
0.76 |
0.75 |
|||
|
4 |
0.80 |
0.70 |
0.75 |
0.74 |
|||
|
5 |
0.67 |
0.64 |
0.65 |
0.74 |
|||
|
Naïve Bayes |
1 |
0.69 |
0.74 |
0.71 |
0.77 |
0.65 |
0.66 |
|
2 |
0.64 |
0.68 |
0.66 |
0.50 |
|||
|
3 |
0.75 |
0.72 |
0.73 |
0.70 |
|||
|
4 |
0.70 |
0.65 |
0.67 |
0.70 |
|||
|
5 |
0.73 |
0.76 |
0.74 |
0.64 |
|||
|
Gradient Boosting |
1 |
0.86 |
0.89 |
0.87 |
0.95 |
0.83 |
0.87 |
|
2 |
0.82 |
0.85 |
0.83 |
0.95 |
|||
|
3 |
0.79 |
0.81 |
0.80 |
0.79 |
|||
|
4 |
0.76 |
0.74 |
0.75 |
0.65 |
|||
|
5 |
0.74 |
0.70 |
0.72 |
0.92 |
The results in table 6 show that the Random Forest and Gradient Boosting models perform best with accuracies of 0.85 and 0.83 respectively, achieving weighted Kappa scores of 0.86 and 0.87, indicating excellent agreement beyond chance for ordinal GMFCS classification, followed by the Decision Tree with an accuracy of 0.81 and a weighted Kappa of 0.80. Random Forest demonstrates exceptional discriminative performance with AUC values ranging from 0.76 to 0.96 across GMFCS levels, with particularly strong performance for levels 1 (AUC=0.94) and 2 (AUC=0.96). Similarly, Gradient Boosting achieves outstanding AUC scores of 0.95 for both levels 1 and 2, demonstrating superior ability to distinguish between mild and moderate severity cases. These models show stability and high performance across the different GMFCS levels, with precision and recall scores that remain high for the majority of classes. Notably, both ensemble methods maintain balanced performance across all severity levels, with Random Forest achieving consistent F1-scores above 0.76 and Gradient Boosting maintaining F1-scores above 0.72 for all GMFCS levels. The SVM, KNN and Naive Bayes models performed less well, with accuracies ranging from 0.65 to 0.70, and correspondingly lower weighted Kappa values (0.66-0.76), indicating moderate agreement for clinical classification, and inconsistent performance between classes. SVM shows reasonable discriminative ability with AUC values between 0.77-0.87, but struggles with class-specific precision, particularly for severe cases (Level 4: Precision=0.65). KNN and Naive Bayes demonstrate the most inconsistent performance, with KNN achieving moderate AUC values (0.66-0.75) and Naive Bayes showing particularly poor discrimination for Level 2 cases (AUC=0.50). Logistic regression offers moderate performance with an accuracy of 0.75 and weighted Kappa of 0.73, representing acceptable clinical agreement, but cannot match the precision and recall of models based on decision trees and ensembles. The superior performance of ensemble methods
(Random Forest and Gradient Boosting) validates their effectiveness in capturing complex, non-linear relationships inherent in cerebral palsy severity assessment, while their high weighted Kappa scores confirm clinical utility for ordinal GMFCS classification. Integrating ontology into machine learning models can considerably improve the performance and robustness of classification systems. Our ontological framework transforms raw patient observations into semantically enriched features, enabling the ML pipeline to extract deeper clinical insights through logical inference and domain knowledge encoding. This semantic enhancement process automatically generates clinically meaningful attributes that capture complex relationships between symptoms, developmental patterns, and severity indicators. In the context of GMFCS level classification, a well-designed ontology can provide a contextual understanding of the relationships between different clinical features and cerebral palsy severity levels. The ontological reasoning serves as a semantic bridge between clinical observations and machine learning algorithms, systematically encoding expert medical knowledge into computable features that mirror clinical decision-making patterns. This approach enables the ML pipeline to work with features that already incorporate domain expertise rather than attempting to discover these complex clinical relationships from raw data alone. This additional knowledge helps machine learning models to better interpret complex data and capture correlations that might be difficult to learn from numerical data alone. By integrating semantic reasoning, our ontology transforms the feature space from simple clinical measurements into rich representations that reflect clinical understanding of cerebral palsy progression and severity assessment. The ML pipeline benefits from this semantic enhancement by receiving features that encode medical expertise, such as mobility potential indicators, developmental milestone alignments, and severity composite scores. This ontology-driven feature extraction process enables machine learning algorithms to focus on pattern recognition within clinically validated feature spaces, resulting in the significant performance improvements observed across all models, with Random Forest and Gradient Boosting achieving accuracies of 85% and 83% respectively through this semantic enhancement approach.
The process of building a machine learning model based on an ontology involves a sequence of key steps as shown in Fig. 9. It all starts with the collection and cleaning of clinical data, eliminating anomalies and managing missing values.
This data is then semantically enriched through alignment with the concepts of an ontology, enabling new features to be created and meaningful metadata to be added. Once the data has been prepared, the relevant features are selected and, if necessary, transformed for learning. The data is then divided into training and test phase to evaluate the performance of the model. An appropriate machine learning algorithm is selected and trained on the training data. The model is then evaluated on the test set using relevant metrics, and adjustments are made if necessary to optimize hyperparameters or feature selection. The best model is finally selected and deployed in production to generate predictions on new data.
To ensure the model’s longevity, continuous monitoring is put in place, and regular updates are made to integrate new data and evolve the ontology if necessary. This approach combines machine learning and ontology to produce models that are more accurate, interpretable and adapted to specific domains.
Our ontology-driven framework enriched the feature gap by embedding clinical semantics into the machine learning pipeline, enabling the model to capture complex relationships between symptoms, diagnoses, and functional outcomes. This semantic enhancement operates through a dual mechanism: first, by transforming discrete clinical observations into contextually aware features that encode medical expertise, and second, by providing inference rules that validate predictions against established clinical knowledge.
Automatically, our results reveal why ensemble methods (Random Forest and Gradient Boosting) benefited more substantially from ontological enhancement than linear models. The ontology generates multiple semantic perspectives on the same clinically available data, developmental milestone alignments, and intervention urgency indicators, creating the feature diversity that ensemble methods require for optimal performance. In contrast, linear models like logistic regression showed modest improvements because they cannot fully exploit the non-linear semantic relationships encoded in our ontological framework.
This semantic enhancement contributed to improved accuracy and interpretability, aligning with findings in ontology highlighting that ontology integration, though beneficial, is not a standalone solution. However, our findings both confirm and extend existing literature in important ways. While studies like El Massari et al., demonstrated ontology-ML integration benefits in diabetes prediction, our work reveals domain-specific challenges in pediatric rehabilitation.
The complexity of cerebral palsy, the use of synthetic data, and the limited input modalities likely constrained further performance improvements. In particular, our results are comparable to recent deep learning models applied to infant movement data, which achieved similar predictive metrics without semantic structuring. However, unlike those approaches, our method offers transparent, explainable predictions grounded in clinical knowledge for real-world application. This comparison reveals a critical trade-off in medical AI: while deep learning approaches may achieve marginally higher accuracy through pattern recognition in high dimensional data, they sacrifice the interpretability that clinicians require for diagnostic confidence.
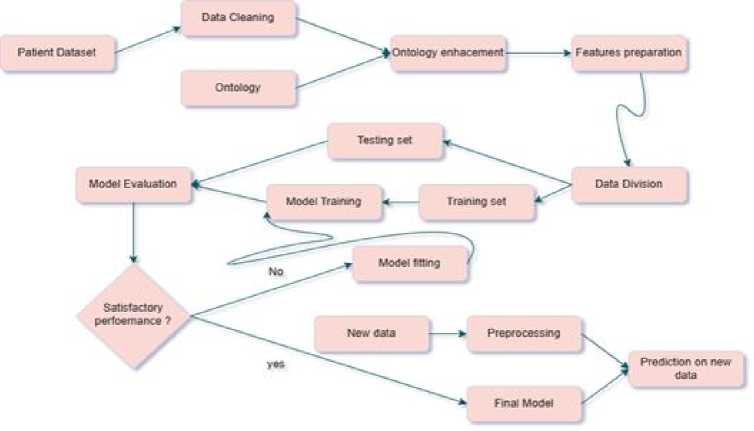
Fig.9. The hierarchical steps of the predictive model
Looking forward, our findings suggest several machine hypotheses for future investigation. First, the superior performance of tree-based models indicates that GMFCS classification follows hierarchical decision patterns that could be explicitly modeled in next generation ontologies. Second, the persistent Levels classification challenge suggests the need for uncertainty quantification mechanisms that can flag ambiguous cases for expert review. Finally, the computational overhead we observed points toward the need for optimized semantic reasoning engines designed specifically for realtime clinical applications.
These findings suggest that integrating ontologies into predictive models is a valuable strategy for early CP severity estimation and personalization of rehabilitation, while also emphasizing the need for richer data, validation, and continuous ontology refinement. However, our work also reveals important limitations of current hybrid approaches: the knowledge intensive ontology development process, the computational overhead, and the challenge of maintaining semantic consistency as medical knowledge evolves. These findings argue for a research agenda focused on automated ontology construction, efficient semantic reasoning algorithms, and standardized frameworks for medical knowledge representation that can evolve with clinical practice.
While most contributors are institutionally based in Tunisia, we collaborated with an Egyptian researcher to contextualize and validate the ontology within the Egyptian healthcare system. This international collaboration reflects our commitment to developing globally relevant and adaptable clinical AI solutions.
6. Conclusions
This study successfully developed OntoML-CP, a novel hybrid intelligence framework that combines machine learning with ontology-based reasoning to predict GMFCS levels in children with cerebral palsy. The integration of OWLbased cerebral palsy ontology with FOAF ontology enabled semantic feature engineering that transformed raw clinical observations into clinically meaningful representations. Random Forest and Gradient Boosting models achieved superior performance with 85% and 83% accuracy, respectively, when enhanced with ontological features, significantly outperforming baseline models (72% to 85% improvement for Random Forest). Beyond achieving strong predictive accuracy, the semantic layer provides transparent, clinically interpretable explanations that link predictions to established medical knowledge, addressing the critical need for explainable AI in pediatric healthcare. This hybrid approach establishes a new paradigm for medical AI that balances computational performance with clinical interpretability, offering healthcare providers a reliable tool for early cerebral palsy severity assessment and personalized rehabilitation pathway planning.
Author Contributions Statement
Rahma Haouas Zahwanie - Conceptualization, Methodology, and Software Implementation: Proposed the research ideas, designed the OntoML-CP framework architecture, developed and implemented the ontological model and machine learning pipeline, and supervised the overall project execution.
Lilia Cheniti-Belcadhi - Review and Editing, and Project Management: Reviewed and edited the manuscript, ensured scientific clarity and coherence, coordinated project milestones, and provided methodological supervision throughout the research process.
Saoussen Layouni - Data Curation and Clinical Validation: Contributed clinical expertise in physical medicine and rehabilitation, supervised the generation and validation of the synthetic dataset based on real clinical cases, and ensured the clinical relevance of the GMFCS classification framework.
Ghada El Khayat - Formal Analysis, Visualization, and Statistical Analysis: Contributed to the formal evaluation of experimental results, reviewed performance metrics and statistical analysis, and supported the validation and contextualization of the ontology within the international healthcare context.
All authors have read and agreed to the published version of the manuscript.
Conflict of Interest Statement
The authors declare no conflicts of interest.
Funding Declaration
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Data Availability Statement
The dataset used in this study is a synthetic dataset representing children with cerebral palsy, generated by physical medicine and rehabilitation specialists based on real clinical cases from the Physical Medicine and Rehabilitation Department at University Hospital center Sahloul, Sousse, Tunisia, and validated through expert review. The dataset is not publicly available at this time but may be made available upon reasonable request to the corresponding author.
Ethical Declarations
The clinical knowledge underlying the synthetic dataset was derived from real patient cases followed at the Physical Medicine and Rehabilitation Department of University Hospital center Sahloul, Sousse, Tunisia. The synthetic dataset was generated and validated by medical specialists to preserve patient anonymity and confidentiality. No direct human subjects were enrolled in this study. All clinical data used as a basis for dataset generation was handled in accordance with institutional ethical guidelines and applicable data protection regulations.
Acknowledgments
The authors sincerely thank the medical specialists at the Physical Medicine and Rehabilitation Department of University Hospital center Sahloul (Sousse, Tunisia) for their professional contribution to the generation and validation of the clinical dataset, and for their valuable expertise in cerebral palsy assessment. We also thank the reviewers for their constructive feedback, which has contributed to improving the quality and reliability of this work.
Declaration of Generative AI in Scholarly Writing
During the preparation of this manuscript, the authors used AI-assisted tools for language editing and grammar checking. After using these tools, the authors reviewed and edited the content as necessary and take full responsibility for the final version of the manuscript.
Abbreviations
The following abbreviations are used in this manuscript:
AI - Artificial Intelligence
AUC - Area Under the Curve
BFO - Basic Formal Ontology
CN - Convolutional Neural Network
CP - Cerebral Palsy
DL - Deep Learning
EHR - Electronic Health Record
FOAF - Friend of a Friend
GB - Gradient Boosting
GMFCS Gross Motor Function Classification System
KNN - K-Nearest Neighbors
LOINC Logical Observation Identifiers Names and Codes
LR - Logistic Regression
ML - Machine Learning
MRI - Magnetic Resonance Imaging
NLP - Natural Language Processing
OntoML-CP - Ontology-driven Machine Learning for Cerebral Palsy
OWL - Web Ontology Language
RDF - Resource Description Framework
RF - Random Forest
RNN - Recurrent Neural Network
SNOMED-CT - Systematized Nomenclature of Medicine - Clinical Terms
SPARQL - SPARQL Protocol and RDF Query Language
SVM - Support Vector Machine
SWRL - Semantic Web Rule Language
UMLS - Unified Medical Language System


