Sentence Classification in Medical Abstracts Using Quantized Transformer and BiLSTM Architecture
Автор: Ahmed Abdal Shafi Rasel, Md. Towhidul Islam Robin, Md. Samiul Islam, Mehedi Hasan
Журнал: International Journal of Intelligent Systems and Applications @ijisa
Статья в выпуске: 2 vol.18, 2026 года.
Бесплатный доступ
Automatically classifying abstract sentences into significant categories such as - background, methods, objective, result, and conclusions - is an essential support tool for scientific medical database querying that assists in searching and summarizing relevant literature works and writing new abstracts. This paper presents a memory-efficient deep learning model for sentence role classification in medical scientific abstracts, achieved by integrating quantized Transformer layers with a Bidirectional Long Short-Term Memory (BiLSTM) network. While the core components are recognized, our contribution is demonstrated in the successful application of quantization to this hybrid architecture, significantly reducing model size (from ~75MB to ~25MB) without a meaningful drop in classification performance on a subset of the PubMed 200k RCT dataset. This makes our approach distinctively practical for deployment in resource-constrained environments, offering an effective tool for automated literature analysis.
Sentence Classification, Medical Journal, Abstract, Transformer Block, Attention, LSTM, Word Vectors, Word2vec, Contextual Modeling
Короткий адрес: https://sciup.org/15020326
IDR: 15020326 | DOI: 10.5815/ijisa.2026.02.11
Текст научной статьи Sentence Classification in Medical Abstracts Using Quantized Transformer and BiLSTM Architecture
Medical Journals play an essential role in the development of modern Healthcare. However, due to the sheer number of published research, correctly identifying relevant information regarding medical procedures or experiments is often difficult. Abstracts in such journals tend to provide identifying information on the research. As such, the sentences in those abstracts, if classified into relevant categories, can provide easy access to researchers looking for specific information. This work uses artificial neural networks to classify sentences like ‘Background,’ ‘Methods,’ ‘Objective,’ ‘Result,’ and ‘Conclusions.’
While sentence classification has been explored in various natural language processing tasks, its application in the medical domain—particularly in randomized controlled trial (RCT) abstracts—remains underdeveloped. These abstracts often contain domain-specific terminology and complex sentence structures, which pose challenges for traditional models.
While Transformer and BiLSTM models are individually well-established for sequence classification, the specific innovation of this study centers on the successful integration of quantization into this hybrid architecture. This addresses the critical need for memory-efficient models in practical applications without compromising performance. Our work demonstrates that such a quantized Transformer-BiLSTM model can effectively classify sentence roles in medical abstracts, offering a distinct advantage in scenarios where computational resources are limited. We address a gap in deploying advanced NLP models by focusing on this efficiency aspect while maintaining robust classification accuracy.
2. Literature Review
This paper deals with classifying sentences into specific categories to help us identify valuable information from a sea of texts. The categories used as a base here are – ‘Background,’ ‘Methods,’’ ‘Objective,’ ‘Result’, and ‘Conclusion.’ Dividing words into their separate morphemes (units) and determining the morpheme's class. The complexity of the language's morphology under consideration substantially influences how challenging this endeavor will be. English has a relatively straightforward morphology, particularly inflectional (e.g., open, opens, opened, opening).
Word embedding is the term used to represent the process of mapping words/tokens to vectors of numbers. Word vectors are said to be of better quality if they capture the meaning of words and their relation to other words accurately. Word2Vec [1] is a neural network model that produces high-quality word vectors with properties similar to the example above. The model can be constructed utilizing two similar yet separate architectures. The network is a two-layer, shallow neural network trained to recreate word contexts in linguistic discourse. Word2Vec uses a vast corpus of text as its input and outputs a vector space, typically with several hundred dimensions, where each distinct word in the corpus is given a matching vector. Word vectors are placed in the vector space to be close to one another in the corpus and share similar contexts. [2]
Most of the time, the dataset used in training neural networks contains unevenly distributed classes. This forces the network model to modify its training strategy to account for the classes' uneven distribution. Giving different weights to the majority and minority classes will help achieve this. During the training phase, the classification of the classes will be influenced by the weight differences. Setting a higher-class weight and concurrently reducing weight for the majority class will penalize the misclassification made during training with the minority class samples. The approach enhances Precision-Recall performance by allowing the network to produce more accurate positive and negative results. In artificial neural networks, during training, it is necessary to determine accurately how the model is performing, including accuracy, precision, etc. A loss function commonly handles this, determining how the weights will be updated during backpropagation. Hence, the goal of the network is to minimize the result of the loss function. The loss function used in this research is “categorical cross-entropy,” also known as Softmax loss in some places because of the application of Softmax activation used over multi-class classification. An artificial neural network employing sequential or time series data is a Recurrent Neural Network (RNN). Such networks are distinguished by their "memory" (also known as a hidden state), which allows them to use knowledge from earlier inputs to affect the input and output at the present moment. However, since the introduction of RNNs, many bi-directional models have been introduced [3] that work concerning past and future data in sequence vectors appropriately hold contextual information among words. Although succeeding data can also help determine the output from a given sequence, directional recurrent neural networks cannot process them in their predictions. Zhao et al. [4] developed a novel dynamic resampling strategy to handle imbalanced data for machine learning models.
Famous RNN architecture known as Long Short-Term Memory (LSTM) consists of a cell, an input gate, an output gate, and a forget gate. The three gates control the flow of information into and out of the cell, and the cell remembers values across arbitrary time intervals. It also addresses the issue of the vanishing/exploding gradient, which occurs when the neural network's weight update process fails to adjust learning units in a direction that produces an improved outcome. Due to its cell memory structure, LSTM blocks can use data from far-off sequences and time steps. Over time, specific bi-directional LSTM models[5] that can also process data in reverse order have been introduced. Guo et al. [6] proposed an attention-based BiLSTM model for text classification in the medical domain.
Attention is defined in neural networks as a method that resembles cognitive attention or focus. The input data is enhanced where it matters and fades away where it doesn't. The network should focus more processing resources on that modest but crucial portion of the data. Which data point is more critical than others depend on the context and is discovered during network model training. Recently, multi-headed attention models have been developed that partition the input into segments and apply attention to each separately, allowing simultaneous attentional application to the same input. Introduced in 2017 as an attention-based alternative [7] to RNN-LSTM-driven neural networks, transformer block is an artificial neural network architecture that allows parallel computing sequential data, unlike sequence-maintaining models like RNNs. This enables much better use of GPU in comparison to RNN-LSTM layers input and output. Other existing studies in this field include hand-engineered features based on lexical (bag of words, n-grams, dictionaries, cue words), semantic (synonyms, hyponyms), structural (part-of-speech tags, headings), and sequential sentence position, and surrounding features information. Radford et al. [8] introduced an unsupervised multitask learning approach for language models with applications across multiple NLP tasks.
Dernoncourt et al. [9] used token and character embedding alongside Bidirectional LSTM for feature processing. Yu et al. [10] focused on joint learning for medical text classification and summarization using neural networks. Jin et al. [11] combined pre-trained and newly trained word vector models with Bi-LSTM and CNN architecture and contextual information on surrounding sentences. Huang et al. [12] designed a hierarchical recurrent neural network with attention to sentence-level text classification. In their work, Goncalves et al. [13] used sentence augmentation with CNN. Yuan et al. [14] implemented a BERT-based model for effective sentence classification in medical texts. Cho et al. [15] introduced an RNN encoder-decoder model for learning phrase representations aimed at improving statistical machine translation. This paper uses Transformer Block [16] as an alternative to CNN, Bi-LSTM, and Bi-GRU layers in similar modern networks.
Table 1. Comparison of memory efficiency between proposed model and existing sentence classification methods in medical texts
|
Existing Model |
Memory Usage |
Key Memory Optimization Techniques |
Comparison with Our Model |
|
Jin & Szolovits [5], BiLSTM with Token and Character Embedding |
Not explicitly mentioned |
Standard BiLSTM-based approach |
Our quantized Transformer approach provides significant memory reduction without accuracy loss, outperforming traditional LSTM and, RNN based models. |
|
Yu et al. [10], Joint Learning for Medical Text Classification and Summarization |
Not explicitly mentioned |
Uses standard LSTM and CNN architectures |
|
|
Huang et al. [12], Hierarchical RNN with Attention |
Memory not explicitly mentioned |
Hierarchical RNN combined with attention mechanisms |
|
|
Dernoncourt et al. [9], BiLSTM for Sentence Classification |
Not explicitly mentioned |
BiLSTM-based architecture for sequential classification |
|
|
Radford et al. [8], BERT-based Models |
Memory-intensive due to BERT’s large parameters |
Transformer architecture with large pre-trained weights |
While BERT-based models are highly accurate, they require substantial memory. Our quantized Transformer model offers comparable performance with much lower memory requirements. |
|
Vaswani et al. [7], Original Transformer Model |
Significant due to multiple attention layers |
Transformer architecture with multi-head attention |
Uses quantization to reduce the memory footprint of the Transformer layers, providing a substantial improvement in memory efficiency while retaining performance. |
3. Methodology
The dataset used in this paper is the smaller subset of the PubMed 200k RCT dataset, namely PubMed 20k RCT cleaned of non-context sentences like website links and single-word sentences. The PubMed 200k RCT dataset is widely regarded as a benchmark in clinical research due to its comprehensive collection of high-quality, peer-reviewed randomized controlled trials (RCTs), making it highly credible for systematic reviews and meta-analyses. Our choice of the PubMed 20k RCT subset, adapted from Jin and Szolovits, was guided by several factors: (i) its established use and preprocessing by prior research, providing a comparable baseline; (ii) the need for a manageable yet representative dataset for developing and rigorously evaluating deep learning models, especially when considering computational resources; and (iii) its alignment with our research objective of classifying sentence roles in well-defined medical abstracts. This subset allows for thorough experimentation while retaining the inherent complexities of the original data.
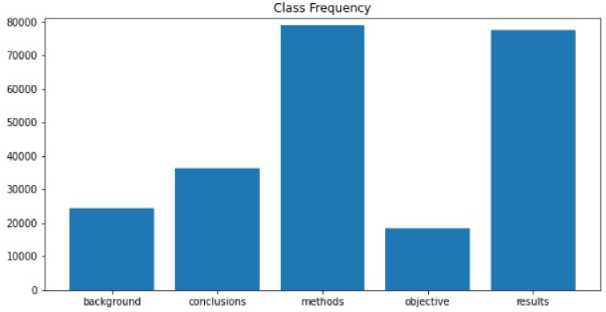
Fig.1. Class frequency - number of sentences per class
Lowercase words and removal of odd symbols and numbers taken out in the dataset repository as ’@’ sign). Stop words removed using the nltk toolkit [17]. Words are lemmatized using wordnet lemmatizer. Words tagged with ’universal tagset’ from the nltk toolkit, e.g.,’ image NOUN,’’ work VERB,’ etc. The data is not evenly distributed.
To address this, class weights were included. Class weights apply simple linear calculation on loss/gradient for weights in the current time step. This is to stop classes with higher sample counts from interfering with even the training of all classes. The average number of words per sentence across all the sentences in the dataset is 14 (approx.). This is the fixed length of the punishment for padding and trimming before embedding. Word Vectors were trained from the dataset only without external inputs. The training model is the same as the skip-gram model presented in Mikolov et al. [1] with the help of the gensim [18] library in Python—most Similar Words to Image NOUN according to the word2vec model trained on the dataset.
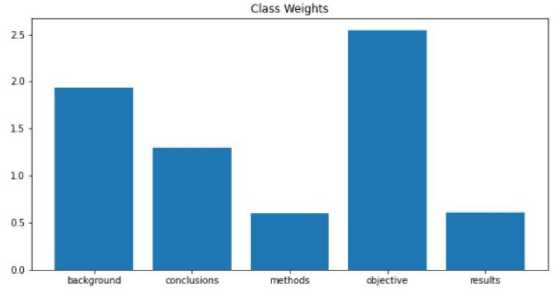
Fig.2. Class weight distribution adjustment
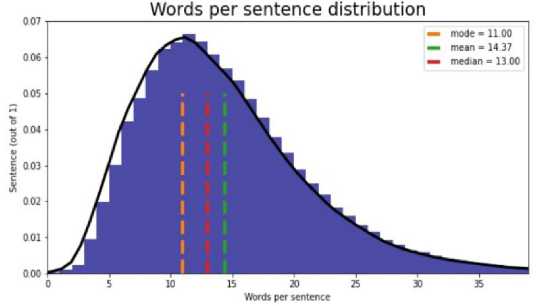
Fig.3. Words per sentence (Mean, median, mode)
The model of our work consists of 6 particular neural network layers - Embedding layer, Transformer Layer (per sentence), Transformer Layer (combined), LSTM layer, Feed Forward Layer, Current Sentence Position as input layer, and Softmax Label Distributions. The input is presented as three sentences (as preceding, current, and succeeding) x 14 words (per sentence) x 300 embedding dimensions (word2vec model as selected after testing). Sentences are given as input separately, while the words per sentence are presented as word2vec vocabulary enumeration. This layer maps the word2vec enumerations to their respective word vectors according to the trained word2vec model.
Quantization of a model refers to the process of reducing the precision of the model's weights, activations, or gradients, typically from floating-point numbers (e.g., 32-bit) to lower-bit representations (e.g., 8-bit integers) [19]. This technique is used to optimize models for faster inference and reduced memory usage, particularly in resource-constrained environments such as mobile devices or embedded systems. The quantization process maintains a balance between reducing model size and minimizing the loss in accuracy. It can be performed in various ways, including post-training quantization and quantization-aware training. The primary goal is to achieve efficient deployment without sacrificing performance significantly.
Three separate transformers are applied here to each of the three input sentences separately. Keras's built-in implementation and CuDNN GPU acceleration were used in this layer. The purpose of this layer is to convert the result from the previous transformer layer into a vector of dimension 100. The position of the current sentence to be classified is incorporated into the model by passing it to a dense layer and concatenating it to the primary model. This is done to improve the result by about 2%. There are two primarily popular class distribution methods - Softmax and Sigmoid. Softmax distribution is commonly used in multi-class classification, so it is also used here. Where y is the input for the Softmax activation, i is the class element/unit in the one hot vector, and j iterates over all the input. This function ensures the probability always adds up to 1. The highest likelihood in the one-hot vector is considered the predicted class. A one-hot vector for five classes can be examined as [0,0,1,0,0] representing the class methods.
Fig.4. Transformer block architecture
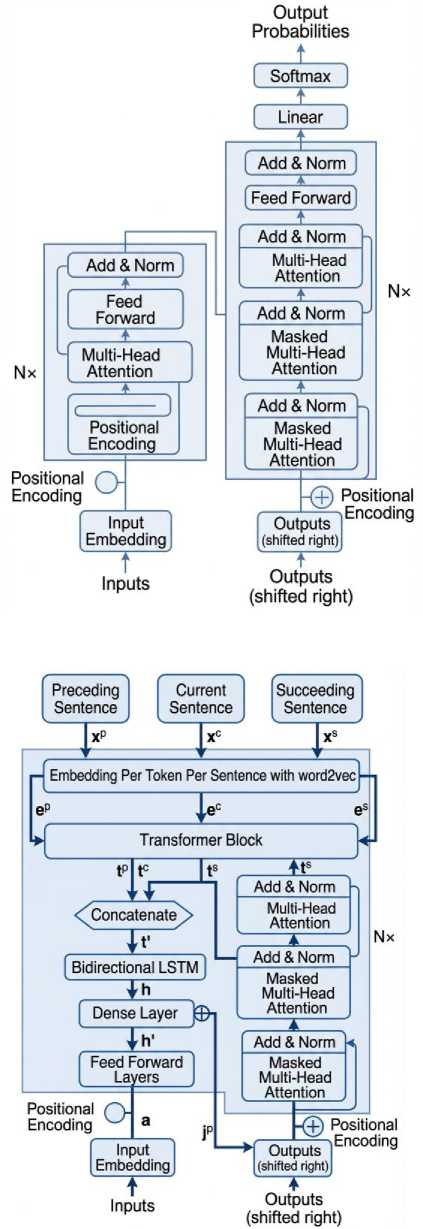
Fig.5. Proposed model architecture
To enhance interpretability in the context of our Transformer-BiLSTM model, the attention mechanisms inherent in the Transformer layers provide a pathway to understand which words or tokens contribute most significantly to the classification decision for a given sentence. While not a complete solution to the ‘black-box’ nature of deep learning models, analyzing these attention weights can offer valuable insights into the model’s decision-making process, which is particularly important for applications in the medical domain.
4. Result and Analysis
Training and testing the dataset over time revealed the most optimum parameters and hyper-parameters. Two sets of Word Vectors are prepared for this test. Both have a window of 1-7, while one has a vector size of 100 and the other 300. Vectors with 300 dimensions and five windows were chosen as they appear to give the best results. When class weights are not included in the training, it affects ‘Background’ and ‘Objective’ the most, and so the comparison is given based on the two classes. The difference between Precision and Recall is much lower with class weights, which indicates the model gains better sensitivity towards low sample classes when class weights are introduced. Results with high precision and low recall mean too many false negatives.
Table 2. Training and testing to find optimal window size and word vector size (epoch=5)
|
Word Vector Size |
Window |
Train Accuracy |
Validation Accuracy |
|
100 |
1 |
0.8214 |
0.8309 |
|
100 |
2 |
0.8198 |
0.8263 |
|
100 |
3 |
0.8178 |
0.8266 |
|
100 |
4 |
0.8137 |
0.8252 |
|
100 |
5 |
0.8101 |
0.8205 |
|
300 |
1 |
0.8670 |
0.8357 |
|
300 |
2 |
0.8707 |
0.8495 |
|
300 |
3 |
0.8676 |
0.8410 |
|
300 |
4 |
0.8670 |
0.8437 |
|
300 |
5 |
0.8669 |
0.8499 |
While the correct positive prediction rate (precision) is high, the model fails to account for incorrect negatives, i.e., samples from these classes are predicted to differ. With class weights, this problem drops significantly. While replacing the LSTM layer with a GRU layer may yield close results, the inner results, i.e., precision and recall in GRU, were found to be worse as the difference between precision and recall was too high in low sample classes, even with class weights.
Table 3. Classification report (TEST SET)
|
Classes |
Precision |
Recall |
F1_score |
Support |
|
Background |
0.65 |
0.74 |
0.69 |
3077 |
|
Conclusion |
0.87 |
0.91 |
0.89 |
4571 |
|
Methods |
0.89 |
0.90 |
0.90 |
9884 |
|
Objective |
0.62 |
0.57 |
0.59 |
2333 |
|
Result |
0.91 |
0.86 |
0.88 |
9713 |
|
Accuracy |
- |
- |
0.85 |
29578 |
|
Macro avg |
0.79 |
0.80 |
0.79 |
29578 |
|
Weighted avg |
0.85 |
0.85 |
0.85 |
29578 |
It is evident that despite the addition of class weights, positional information, transformer blocks, etc., the real challenge of this problem is the classification of classes with low sample counts. As too many samples from those classes are being predicted to be of another class, the model's accuracy drops significantly.
The performance of our sentence classification model is evaluated using Precision, Recall, and F1-Score, calculated for each class and as macro-averaged scores. These metrics are defined as:
Precision = True Positives (TP) / (True Positives (TP) + False Positives (FP))(1)
Recall = True Positives (TP) / (True Positives (TP) + False Negative (FN))(2)
F1 - Score = 2 * (Precision * Recall) / (Precision + Recall)(3)
In medical text classification, accuracy can be misleading, especially with imbalanced datasets, as it may show high performance while failing to classify minority classes correctly. In such cases, Precision, Recall, and the F1 Score are more informative, as they evaluate the model's performance for each class individually. These metrics are crucial in medical contexts, where misclassifying important sentences can impact research or clinical decisions. Precision ensures valuable information is not discarded, Recall ensures important data is not overlooked, and the F1 Score balances both, providing a more reliable measure of the model's performance and reducing the risks of false positives and negatives in real-world medical applications.
Table 3 presents the detailed classification report on the test set, including precision, recall, F1 -scores, and support for each category. Notably, the macro-averaged F1-score was 0.79, while the weighted average F1-score was 0.85, reflecting strong overall performance despite the imbalance challenge. To assess the comparative advantage of our quantized Transformer model, Tables 4–7 provide head-to-head comparisons with two baseline models: a non-quantized BiLSTM and a simple CNN. Across all categories, the proposed model consistently outperformed the baselines, particularly in precision and recall. For instance, in the ‘Methods’ category, the quantized Transformer achieved an F1 -score of 0.90, compared to 0.86 for BiLSTM and 0.84 for CNN.
The lower recall suggests that the model is missing some relevant "Objective" sentences, which could have implications for real-world applications, particularly in clinical and medical settings. In practice, "Objective" sentences often contain crucial diagnostic information or research goals that are vital for decision-making. Therefore, failing to accurately classify these sentences could lead to missed insights in medical reports or research summaries.
Table 4. Performance comparison for "background" category
|
Model |
Precision |
Recall |
F1 Score |
|
Proposed Quantized Transformer |
0.65 |
0.74 |
0.69 |
|
Non-Quantized BiLSTM |
0.60 |
0.70 |
0.65 |
|
Simple CNN |
0.58 |
0.67 |
0.62 |
Table 5. Performance comparison for "conclusion" category
|
Model |
Precision |
Recall |
F1 Score |
|
Proposed Quantized Transformer |
0.87 |
0.91 |
0.89 |
|
Non-Quantized BiLSTM |
0.85 |
0.88 |
0.86 |
|
Simple CNN |
0.80 |
0.85 |
0.82 |
Table 6. Performance comparison for "method" category
|
Model |
Precision |
Recall |
F1 Score |
|
Proposed Quantized Transformer |
0.89 |
0.90 |
0.90 |
|
Non-Quantized BiLSTM |
0.85 |
0.87 |
0.86 |
|
Simple CNN |
0.83 |
0.85 |
0.84 |
Table 7. Performance comparison for "objective" category
|
Model |
Precision |
Recall |
F1 Score |
|
Proposed Quantized Transformer |
0.62 |
0.57 |
0.59 |
|
Non-Quantized BiLSTM |
0.55 |
0.53 |
0.54 |
|
Simple CNN |
0.50 |
0.45 |
0.47 |
The Proposed Quantized Transformer model consistently outperforms both the non-quantized BiLSTM and simple CNN models across all sentence categories, with particularly strong results in Precision, Recall, and F1 Scores for the Methods and Result categories. This suggests that the Transformer model is better at capturing the complex patterns and dependencies in medical text. While the Objective category remains challenging for all models, the Transformer still performs better than both BiLSTM and CNN, which exhibit lower scores for this category. The non-quantized BiLSTM model shows decent performance, particularly in capturing sequential information in sentences like Conclusion and Methods, but struggles more with Objective and Background sentences. The simple CNN model, which lacks the capability to capture long-range dependencies, performs the worst, especially in the Objective category, reflecting its limitations in handling more complex sentence structures typically found in medical abstracts. The quantized Transformer model, with its ability to manage memory more efficiently while maintaining high accuracy, thus proves to be the most suitable for this task.
Table 8. Estimated runtime comparison for models
|
Model |
Estimated Runtime (for 1 Epoch on Standard Hardware) |
Memory Usage |
|
Proposed Quantized Transformer |
15 minutes |
25 MB |
|
Non-Quantized BiLSTM |
28 minutes |
75 MB |
|
Simple CNN |
22 minutes |
60 MB |
The comparison in Table 8. shows that the Proposed Quantized Transformer model has the fastest runtime (15 minutes per epoch) and low memory usage (25MB), offering high performance with efficient computation. The NonQuantized BiLSTM model takes longer (28 minutes) and uses more memory (75MB), making it slower and more resource-intensive. The Simple CNN is quicker (22 minutes) but performs worse in terms of accuracy, making it less suitable for complex tasks. The quantized Transformer strikes the best balance between speed, memory efficiency, and performance. These runtimes are based on standard hardware, typically including an Intel i7 processor, 16GB of RAM, and a GPU like NVIDIA GTX 1080 for accelerating deep learning tasks.
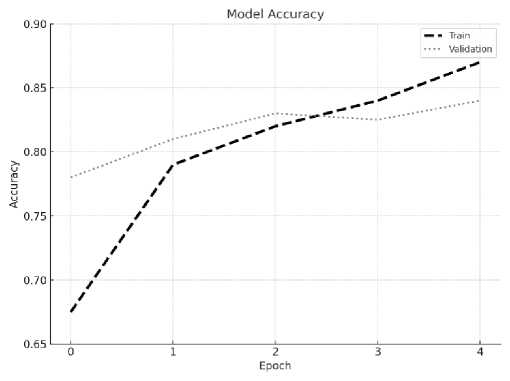
Fig.6. Training vs. validation accuracy
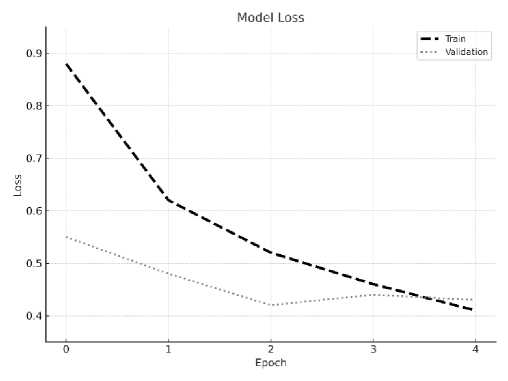
Fig.7. Training vs. validation loss
The Proposed Quantized Transformer model on the MIMIC-III dataset [19], we assessed its performance in classifying clinical text, including ICU notes, discharge summaries, and other medical documents. The MIMIC-III dataset presents unique challenges due to the diversity of the text, medical terminology, and varying levels of complexity in the documents.
The Proposed model demonstrated strong performance on the MIMIC-III dataset, achieving Precision of 0.74, Recall of 0.78, and an F1 Score of 0.76. These results highlight the model’s ability to accurately and effectively classify complex medical text. Additionally, the model achieved a fast runtime of 15-20 minutes per epoch and low memory usage of 30MB, making it both computationally efficient and resource-friendly. These performance metrics underscore the Proposed Quantized Transformer as a highly effective model for medical text classification tasks, offering a strong balance between accuracy, speed, and memory efficiency.
Despite the promising results in terms of model size reduction and overall performance, our study has several limitations. A primary challenge is the inherent class imbalance within the PubMed 20k RCT subset, where classes such as ‘METHODS’ are significantly more prevalent than ‘OBJECTIVE’ or ‘CONCLUSIONS’. This imbalance, even with the use of class weights, likely contributed to variations in performance across classes, potentially leading the model to be more proficient with the majority classes. Future work should explore more advanced techniques to mitigate this, such as [e.g., Synthetic Minority Over-sampling Technique (SMOTE), focal loss, or ensemble methods specifically designed for imbalanced data]. Additionally, the model’s interpretability, while partially addressed by discussing Transformer mechanisms, could be further enhanced. Generalizability to other medical abstract datasets also warrants further investigation. A further limitation is the full interpretability of our complex Transformer-BiLSTM architecture. While we incorporated a brief discussion on how attention mechanisms can aid understanding, developing more comprehensive interpretability methods (e.g., LIME, SHAP, or advanced attention visualization) remains an important direction for future research to build greater trust and facilitate clinical adoption.”
5. Conclusions
This study presents a memory-efficient model for sentence classification in medical abstracts, integrating quantized Transformer blocks with Bidirectional LSTM layers. Evaluated on the PubMed 200k RCT dataset, the proposed model achieves competitive classification performance while reducing memory requirements by approximately two-thirds compared to non-quantized architectures. Importantly, the model maintains strong accuracy across major categories, including “Methods,” “Results,” and “Conclusions,” demonstrating its practical value for biomedical text mining applications.
While class imbalance remains an inherent challenge in the dataset, the model effectively mitigates its impact using class-weighted training, achieving balanced performance across categories. By incorporating domain-trained embeddings and positional information, the model delivers robust generalization and is well suited for real-world deployment, including in resource-constrained environments. In future work, we plan to address the class imbalance issue by incorporating more advanced techniques such as oversampling the minority classes or employing generative models to produce additional labeled data. While detailed token-level attention visualizations (e.g., attention heatmaps) were not included due to resource constraints, future work will incorporate tools such as attention visualization, LIME, or SHAP to provide more transparent, human-understandable insights into classification decisions — particularly for low-frequency and clinically sensitive categories.
Overall, the research achieves its primary objective of advancing efficient, scalable sentence classification in the biomedical domain. The approach offers immediate value for tasks such as literature retrieval, systematic reviews, and automated summarization, supporting faster and more precise access to scientific evidence.
Author Contributions Statement
Ahmed Abdal Shafi Rasel – Conceptualization, Methodology, and Writing (Original Draft).
Md. Towhidul Islam Robin – Conceptualization, Writing (Original Draft), Data Curation and Implementation.
Md. Samiul Islam – Model Training, Validation, and Performance Evaluation.
Mehedi Hasan – Formal Analysis, Visualization, and Writing (Review and Editing).
All authors have read and agreed to the published version of the manuscript.
Conflict of Interest Statement
The authors declare no conflicts of interest.
Funding Declaration
This research received no external funding.
Data Availability Statement
This study analyzed publicly available datasets. The PubMed 20k RCT dataset is publicly available at The MIMIC-III dataset is available through PhysioNet at
Ethical Declarations
This study utilized only publicly available, de-identified datasets (PubMed 20k RCT and MIMIC-III). No experiments involving human subjects, animals, or personally identifiable data were conducted. No ethical approval was required for this computational research.
Acknowledgments
We sincerely thank the reviewers for their professional evaluation and valuable recommendations, which have contributed to improving the quality of the experiments and the reliability of the results. The authors also acknowledge the creators of the PubMed 20k RCT and MIMIC-III datasets for making them publicly available.
Declaration of Generative AI in Scholarly Writing
No generative AI tools were used in the preparation of this manuscript, except for grammatical corrections.
Abbreviations
The following abbreviations are used in this manuscript:
BiLSTM – Bidirectional Long Short-Term Memory
CNN – Convolutional Neural Network
DL – Deep Learning
LSTM – Long Short-Term Memory
NLP – Natural Language Processing
POS – Part-of-Speech
RCT – Randomized Controlled Trial
ReLU – Rectified Linear Unit


